Poison Exon Splicing Regulates a Coordinated Network of SR Protein Expression during Differentiation and Tumorigenesis
- PMID: 33176162
- PMCID: PMC7680420
- DOI: 10.1016/j.molcel.2020.10.019
Poison Exon Splicing Regulates a Coordinated Network of SR Protein Expression during Differentiation and Tumorigenesis
Abstract
The RNA isoform repertoire is regulated by splicing factor (SF) expression, and alterations in SF levels are associated with disease. SFs contain ultraconserved poison exon (PE) sequences that exhibit greater identity across species than nearby coding exons, but their physiological role and molecular regulation is incompletely understood. We show that PEs in serine-arginine-rich (SR) proteins, a family of 14 essential SFs, are differentially spliced during induced pluripotent stem cell (iPSC) differentiation and in tumors versus normal tissues. We uncover an extensive cross-regulatory network of SR proteins controlling their expression via alternative splicing coupled to nonsense-mediated decay. We define sequences that regulate PE inclusion and protein expression of the oncogenic SF TRA2β using an RNA-targeting CRISPR screen. We demonstrate location dependency of RS domain activity on regulation of TRA2β-PE using CRISPR artificial SFs. Finally, we develop splice-switching antisense oligonucleotides to reverse the increased skipping of TRA2β-PE detected in breast tumors, altering breast cancer cell viability, proliferation, and migration.
Keywords: RNA splicing, SR proteins, differentiation, cancer, cross-regulation, antisense oligonucleotides, CRISPR/Cas13, CRISPR-Artificial Splicing Factors, alternative splicing, oncogene.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests A patent application concerning this work is in preparation.
Figures

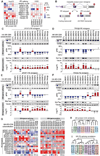





Similar articles
-
Antisense oligonucleotide-mediated TRA2β poison exon inclusion induces the expression of a lncRNA with anti-tumor effects.Nat Commun. 2025 Feb 15;16(1):1670. doi: 10.1038/s41467-025-56913-8. Nat Commun. 2025. PMID: 39955311 Free PMC article.
-
An ultra-conserved poison exon in the Tra2b gene encoding a splicing activator is essential for male fertility and meiotic cell division.EMBO J. 2025 Feb;44(3):877-902. doi: 10.1038/s44318-024-00344-6. Epub 2025 Jan 2. EMBO J. 2025. PMID: 39748121 Free PMC article.
-
Autoregulated splicing of TRA2β programs T cell fate in response to antigen-receptor stimulation.Science. 2024 Sep 13;385(6714):eadj1979. doi: 10.1126/science.adj1979. Epub 2024 Sep 13. Science. 2024. PMID: 39265028 Free PMC article.
-
How does Tra2β protein regulate tissue-specific RNA splicing?Biochem Soc Trans. 2012 Aug;40(4):784-8. doi: 10.1042/BST20120036. Biochem Soc Trans. 2012. PMID: 22817734 Free PMC article. Review.
-
Serine/Arginine-rich protein family of splicing regulators: New approaches to study splice isoform functions.Plant Sci. 2019 Jun;283:127-134. doi: 10.1016/j.plantsci.2019.02.017. Epub 2019 Mar 12. Plant Sci. 2019. PMID: 31128682 Review.
Cited by
-
Zinc supplementation prevents arsenic-induced dysregulation of ZRANB2 splice function.Environ Toxicol Pharmacol. 2022 Aug;94:103921. doi: 10.1016/j.etap.2022.103921. Epub 2022 Jun 25. Environ Toxicol Pharmacol. 2022. PMID: 35764259 Free PMC article.
-
Exploring the Diverse Functional and Regulatory Consequences of Alternative Splicing in Development and Disease.Front Genet. 2021 Nov 24;12:775395. doi: 10.3389/fgene.2021.775395. eCollection 2021. Front Genet. 2021. PMID: 34899861 Free PMC article. Review.
-
Splicing control by PHF5A is crucial for melanoma cell survival.Cell Prolif. 2025 Feb;58(2):e13741. doi: 10.1111/cpr.13741. Epub 2024 Aug 30. Cell Prolif. 2025. PMID: 39212334 Free PMC article.
-
Incomplete paralog compensation generates selective dependency on TRA2A in cancer.PLoS Genet. 2025 May 14;21(5):e1011685. doi: 10.1371/journal.pgen.1011685. eCollection 2025 May. PLoS Genet. 2025. PMID: 40367120 Free PMC article.
-
Thermoregulated transcriptomics: the molecular basis and biological significance of temperature-dependent alternative splicing.Biochem J. 2024 Aug 7;481(15):999-1013. doi: 10.1042/BCJ20230410. Biochem J. 2024. PMID: 39083035 Free PMC article. Review.
References
-
- Ajiro M, Tang S, Doorbar J, and Zheng ZM (2016). Serine/Arginine-Rich Splicing Factor 3 and Heterogeneous Nuclear Ribonucleoprotein A1 Regulate Alternative RNA Splicing and Gene Expression of Human Papillomavirus 18 through Two Functionally Distinguishable cis Elements. J Virol 90, 9138–9152. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

