The Iron-Sulfur Flavoprotein DsrL as NAD(P)H:Acceptor Oxidoreductase in Oxidative and Reductive Dissimilatory Sulfur Metabolism
- PMID: 33178160
- PMCID: PMC7596348
- DOI: 10.3389/fmicb.2020.578209
The Iron-Sulfur Flavoprotein DsrL as NAD(P)H:Acceptor Oxidoreductase in Oxidative and Reductive Dissimilatory Sulfur Metabolism
Erratum in
-
Erratum: The Iron-Sulfur Flavoprotein DsrL as NAD(P)H:Acceptor Oxidoreductase in Oxidative and Reductive Dissimilatory Sulfur Metabolism.Front Microbiol. 2021 Jan 13;11:644616. doi: 10.3389/fmicb.2020.644616. eCollection 2020. Front Microbiol. 2021. PMID: 33519796 Free PMC article.
Abstract
DsrAB-type dissimilatory sulfite reductase is a key enzyme of microbial sulfur-dependent energy metabolism. Sulfur oxidizers also contain DsrL, which is essential for sulfur oxidation in Allochromatium vinosum. This NAD(P)H oxidoreductase acts as physiological partner of oxidative-type rDsrAB. Recent analyses uncovered that DsrL is not confined to sulfur oxidizers but also occurs in (probable) sulfate/sulfur-reducing bacteria. Here, phylogenetic analysis revealed a separation into two major branches, DsrL-1, with two subgroups, and DsrL-2. When present in organisms with reductive-type DsrAB, DsrL is of type 2. In the majority of cases oxidative-type rDsrAB occurs with DsrL-1 but combination with DsrL-2-type enzymes is also observed. Three model DsrL proteins, DsrL-1A and DsrL-1B from the sulfur oxidizers A. vinosum and Chlorobaculum tepidum, respectively, as well as DsrL-2 from thiosulfate- and sulfur-reducing Desulfurella amilsii were kinetically characterized. DaDsrL-2 is active with NADP(H) but not with NAD(H) which we relate to a conserved YRR-motif in the substrate-binding domains of all DsrL-2 enzymes. In contrast, AvDsrL-1A has a strong preference for NAD(H) and the CtDsrL-1B enzyme is completely inactive with NADP(H). Thus, NAD+ as well as NADP+ are suitable in vivo electron acceptors for rDsrABL-1-catalyzed sulfur oxidation, while NADPH is required as electron donor for sulfite reduction. This observation can be related to the lower redox potential of the NADPH/NADP+ than the NADH/NAD+ couple under physiological conditions. Organisms with a rdsrAB and dsrL-1 gene combination can be confidently identified as sulfur oxidizers while predictions for organisms with other combinations require much more caution and additional information sources.
Keywords: DsrAB; DsrL; NAD(P)H; dissimilatory sulfate reduction; dissimilatory sulfur oxidation; sulfite reductase; sulfur metabolism.
Copyright © 2020 Löffler, Wallerang, Venceslau, Pereira and Dahl.
Figures



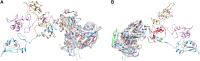
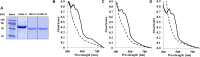
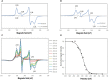

References
LinkOut - more resources
Full Text Sources

