Color Data v2: a user-friendly, open-access database with hereditary cancer and hereditary cardiovascular conditions datasets
- PMID: 33181822
- PMCID: PMC7661094
- DOI: 10.1093/database/baaa083
Color Data v2: a user-friendly, open-access database with hereditary cancer and hereditary cardiovascular conditions datasets
Abstract
Publicly available genetic databases promote data sharing and fuel scientific discoveries for the prevention, treatment and management of disease. In 2018, we built Color Data, a user-friendly, open access database containing genotypic and self-reported phenotypic information from 50 000 individuals who were sequenced for 30 genes associated with hereditary cancer. In a continued effort to promote access to these types of data, we launched Color Data v2, an updated version of the Color Data database. This new release includes additional clinical genetic testing results from more than 18 000 individuals who were sequenced for 30 genes associated with hereditary cardiovascular conditions as well as polygenic risk scores for breast cancer, coronary artery disease and atrial fibrillation. In addition, we used self-reported phenotypic information to implement the following four clinical risk models: Gail Model for 5-year risk of breast cancer, Claus Model for lifetime risk of breast cancer, simple office-based Framingham Coronary Heart Disease Risk Score for 10-year risk of coronary heart disease and CHARGE-AF simple score for 5-year risk of atrial fibrillation. These new features and capabilities are highlighted through two sample queries in the database. We hope that the broad dissemination of these data will help researchers continue to explore genotype-phenotype correlations and identify novel variants for functional analysis, enabling scientific discoveries in the field of population genomics. Database URL: https://data.color.com/.
© The Author(s) 2020. Published by Oxford University Press.
Figures

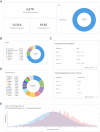
References
-
- Kwon D.H.-M., Borno H.T., Cheng H.H. et al. (2019) Ethnic disparities among men with prostate cancer undergoing germline testing. Urol. Oncol., 38, 80.e1–80.e7. - PubMed
-
- (2018) Science Extension | Garvan institute of medical research https://www.garvan.org.au/research/kinghorn-centre-for-clinical-genomics... (accessed Jan 13, 2020).
-
- Gail M.H., Brinton L.A., Byar D.P. et al. (1989) Projecting individualized probabilities of developing breast cancer for white females who are being examined annually. J. Natl. Cancer Inst., 81, 1879–1886. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

