HIV-1 Gag Forms Ribonucleoprotein Complexes with Unspliced Viral RNA at Transcription Sites
- PMID: 33182496
- PMCID: PMC7696413
- DOI: 10.3390/v12111281
HIV-1 Gag Forms Ribonucleoprotein Complexes with Unspliced Viral RNA at Transcription Sites
Abstract
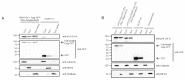
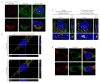
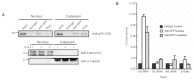
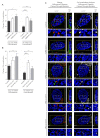
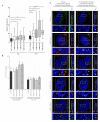
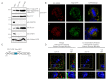
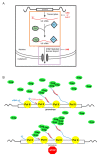
The ability of the retroviral Gag protein of Rous sarcoma virus (RSV) to transiently traffic through the nucleus is well-established and has been implicated in genomic RNA (gRNA) packaging Although other retroviral Gag proteins (human immunodeficiency virus type 1, HIV-1; feline immunodeficiency virus, FIV; Mason-Pfizer monkey virus, MPMV; mouse mammary tumor virus, MMTV; murine leukemia virus, MLV; and prototype foamy virus, PFV) have also been observed in the nucleus, little is known about what, if any, role nuclear trafficking plays in those viruses. In the case of HIV-1, the Gag protein interacts in nucleoli with the regulatory protein Rev, which facilitates nuclear export of gRNA. Based on the knowledge that RSV Gag forms viral ribonucleoprotein (RNPs) complexes with unspliced viral RNA (USvRNA) in the nucleus, we hypothesized that the interaction of HIV-1 Gag with Rev could be mediated through vRNA to form HIV-1 RNPs. Using inducible HIV-1 proviral constructs, we visualized HIV-1 Gag and USvRNA in discrete foci in the nuclei of HeLa cells by confocal microscopy. Two-dimensional co-localization and RNA-immunoprecipitation of fractionated cells revealed that interaction of nuclear HIV-1 Gag with USvRNA was specific. Interestingly, treatment of cells with transcription inhibitors reduced the number of HIV-1 Gag and USvRNA nuclear foci, yet resulted in an increase in the degree of Gag co-localization with USvRNA, suggesting that Gag accumulates on newly synthesized viral transcripts. Three-dimensional imaging analysis revealed that HIV-1 Gag localized to the perichromatin space and associated with USvRNA and Rev in a tripartite RNP complex. To examine a more biologically relevant cell, latently infected CD4+ T cells were treated with prostratin to stimulate NF-κB mediated transcription, demonstrating striking localization of full-length Gag at HIV-1 transcriptional burst site, which was labelled with USvRNA-specific riboprobes. In addition, smaller HIV-1 RNPs were observed in the nuclei of these cells. These data suggest that HIV-1 Gag binds to unspliced viral transcripts produced at the proviral integration site, forming vRNPs in the nucleus.
Keywords: HIV-1 Gag; Rev; nucleus; viral RNA.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Dynamic interactions of retroviral Gag condensates with nascent viral RNA at transcriptional burst sites: implications for genomic RNA packaging.bioRxiv [Preprint]. 2025 Jun 2:2025.01.11.632546. doi: 10.1101/2025.01.11.632546. bioRxiv. 2025. Update in: mBio. 2025 Aug 19:e0169525. doi: 10.1128/mbio.01695-25. PMID: 39829876 Free PMC article. Updated. Preprint.
-
Dynamic interactions of retroviral Gag condensates with nascent viral RNA at transcriptional burst sites: implications for genomic RNA packaging.mBio. 2025 Aug 19:e0169525. doi: 10.1128/mbio.01695-25. Online ahead of print. mBio. 2025. PMID: 40827928
-
Visualizing Association of the Retroviral Gag Protein with Unspliced Viral RNA in the Nucleus.mBio. 2020 Apr 7;11(2):e00524-20. doi: 10.1128/mBio.00524-20. mBio. 2020. PMID: 32265329 Free PMC article.
-
Cessation of HIV-1 transcription by inhibiting regulatory protein Rev-mediated RNA transport.Curr HIV Res. 2009 Jan;7(1):101-8. doi: 10.2174/157016209787048564. Curr HIV Res. 2009. PMID: 19149559 Review.
-
A structurally plastic ribonucleoprotein complex mediates post-transcriptional gene regulation in HIV-1.Wiley Interdiscip Rev RNA. 2016 Jul;7(4):470-86. doi: 10.1002/wrna.1342. Epub 2016 Mar 1. Wiley Interdiscip Rev RNA. 2016. PMID: 26929078 Free PMC article. Review.
Cited by
-
Advances in HIV-1 Assembly.Viruses. 2022 Feb 26;14(3):478. doi: 10.3390/v14030478. Viruses. 2022. PMID: 35336885 Free PMC article. Review.
-
Dynamic interactions of retroviral Gag condensates with nascent viral RNA at transcriptional burst sites: implications for genomic RNA packaging.bioRxiv [Preprint]. 2025 Jun 2:2025.01.11.632546. doi: 10.1101/2025.01.11.632546. bioRxiv. 2025. Update in: mBio. 2025 Aug 19:e0169525. doi: 10.1128/mbio.01695-25. PMID: 39829876 Free PMC article. Updated. Preprint.
-
Role of the Psi Packaging Signal and Dimerization Initiation Sequence in the Organization of Rous Sarcoma Virus Gag-gRNA Co-Condensates.Viruses. 2025 Jan 13;17(1):97. doi: 10.3390/v17010097. Viruses. 2025. PMID: 39861886 Free PMC article.
-
Comparative analysis of retroviral Gag-host cell interactions: focus on the nuclear interactome.bioRxiv [Preprint]. 2024 Mar 6:2024.01.18.575255. doi: 10.1101/2024.01.18.575255. bioRxiv. 2024. Update in: Retrovirology. 2024 Jun 19;21(1):13. doi: 10.1186/s12977-024-00645-y. PMID: 38293010 Free PMC article. Updated. Preprint.
-
Biomolecular condensates: insights into early and late steps of the HIV-1 replication cycle.Retrovirology. 2023 Apr 7;20(1):4. doi: 10.1186/s12977-023-00619-6. Retrovirology. 2023. PMID: 37029379 Free PMC article. Review.
References
-
- Rabson A.B., Graves B.J. Synthesis and Processing of Viral RNA. In: Coffin J.M., Hughes S.H., Varmus H.E., editors. Retroviruses. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY, USA: 1997. - PubMed
-
- Chen J., Rahman S.A., Nikolaitchik O.A., Grunwald D., Sardo L., Burdick R.C., Plisov S., Liang E., Tai S., Pathak V.K., et al. HIV-1 RNA genome dimerizes on the plasma membrane in the presence of Gag protein. Proc. Natl. Acad. Sci. USA. 2016;113:E201–E208. doi: 10.1073/pnas.1518572113. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

