Expansion segments in bacterial and archaeal 5S ribosomal RNAs
- PMID: 33184227
- PMCID: PMC7812874
- DOI: 10.1261/rna.077123.120
Expansion segments in bacterial and archaeal 5S ribosomal RNAs
Abstract
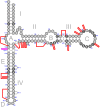
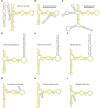
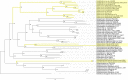
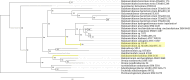
The large ribosomal RNAs of eukaryotes frequently contain expansion sequences that add to the size of the rRNAs but do not affect their overall structural layout and are compatible with major ribosomal function as an mRNA translation machine. The expansion of prokaryotic ribosomal RNAs is much less explored. In order to obtain more insight into the structural variability of these conserved molecules, we herein report the results of a comprehensive search for the expansion sequences in prokaryotic 5S rRNAs. Overall, 89 expanded 5S rRNAs of 15 structural types were identified in 15 archaeal and 36 bacterial genomes. Expansion segments ranging in length from 13 to 109 residues were found to be distributed among 17 insertion sites. The strains harboring the expanded 5S rRNAs belong to the bacterial orders Clostridiales, Halanaerobiales, Thermoanaerobacterales, and Alteromonadales as well as the archael order Halobacterales When several copies of a 5S rRNA gene are present in a genome, the expanded versions may coexist with normal 5S rRNA genes. The insertion sequences are typically capable of forming extended helices, which do not seemingly interfere with folding of the conserved core. The expanded 5S rRNAs have largely been overlooked in 5S rRNA databases.
Keywords: 5S rRNA; archaea; bacteria; expansion segment; ribosome.
© 2021 Stepanov and Fox; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures




Similar articles
-
The evolutionary history of the structure of 5S ribosomal RNA.J Mol Evol. 2009 Nov;69(5):430-43. doi: 10.1007/s00239-009-9264-z. Epub 2009 Jul 29. J Mol Evol. 2009. PMID: 19639237
-
5SRNAdb: an information resource for 5S ribosomal RNAs.Nucleic Acids Res. 2016 Jan 4;44(D1):D180-3. doi: 10.1093/nar/gkv1081. Epub 2015 Oct 20. Nucleic Acids Res. 2016. PMID: 26490961 Free PMC article.
-
Mapping posttranscriptional modifications in 5S ribosomal RNA by MALDI mass spectrometry.RNA. 2000 Feb;6(2):296-306. doi: 10.1017/s1355838200992148. RNA. 2000. PMID: 10688367 Free PMC article.
-
The nucleotide sequence of 5 S ribosomal RNA from Vibrio marinus.Microbiol Sci. 1984 Dec;1(9):229-31. Microbiol Sci. 1984. PMID: 6086080 Review.
-
5S rRNA and ribosome.Biochemistry (Mosc). 2011 Dec;76(13):1450-64. doi: 10.1134/S0006297911130062. Biochemistry (Mosc). 2011. PMID: 22339598 Review.
Cited by
-
A Comparative Perspective on Ribosome Biogenesis: Unity and Diversity Across the Tree of Life.Methods Mol Biol. 2022;2533:3-22. doi: 10.1007/978-1-0716-2501-9_1. Methods Mol Biol. 2022. PMID: 35796979 Free PMC article. Review.
-
Features of smaller ribosomes in candidate phyla radiation (CPR) bacteria revealed with a molecular evolutionary analysis.RNA. 2022 Aug;28(8):1041-1057. doi: 10.1261/rna.079103.122. Epub 2022 Jun 10. RNA. 2022. PMID: 35688647 Free PMC article.
-
Looking through the Lens of the Ribosome Biogenesis Evolutionary History: Possible Implications for Archaeal Phylogeny and Eukaryogenesis.Mol Biol Evol. 2022 Apr 11;39(4):msac054. doi: 10.1093/molbev/msac054. Mol Biol Evol. 2022. PMID: 35275997 Free PMC article.
-
Coevolution of ribosomal RNA expansion segment 7L and assembly factor Noc2p specializes the ribosome biogenesis pathway between Saccharomyces cerevisiae and Candida albicans.Nucleic Acids Res. 2021 May 7;49(8):4655-4667. doi: 10.1093/nar/gkab218. Nucleic Acids Res. 2021. PMID: 33823547 Free PMC article.
-
Ribosome Biogenesis in Archaea.Front Microbiol. 2021 Jul 22;12:686977. doi: 10.3389/fmicb.2021.686977. eCollection 2021. Front Microbiol. 2021. PMID: 34367089 Free PMC article. Review.
References
-
- Armache J-P, Anger AM, Marquez V, Franckenberg S, Fröhlich T, Villa E, Berninghausen O, Thomm M, Arnold GJ, Beckmann R. 2012. Promiscuous behaviour of archaeal ribosomal proteins: implications for eukaryotic ribosome evolution. Nucleic Acids Res 41: 1284–1293. 10.1093/nar/gks1259 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
