A natriuretic peptide from Arabidopsis thaliana (AtPNP-A) can modulate catalase 2 activity
- PMID: 33184368
- PMCID: PMC7665192
- DOI: 10.1038/s41598-020-76676-0
A natriuretic peptide from Arabidopsis thaliana (AtPNP-A) can modulate catalase 2 activity
Abstract
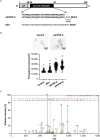
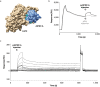
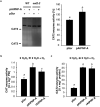
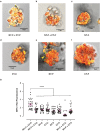
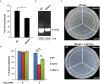
Analogues of vertebrate natriuretic peptides (NPs) present in plants, termed plant natriuretic peptides (PNPs), comprise a novel class of hormones that systemically affect salt and water balance and responses to plant pathogens. Several lines of evidence indicate that Arabidopsis thaliana PNP (AtPNP-A) affects cellular redox homeostasis, which is also typical for the signaling of its vertebrate analogues, but the molecular mechanism(s) of this effect remains elusive. Here we report identification of catalase 2 (CAT2), an antioxidant enzyme, as an interactor of AtPNP-A. The full-length AtPNP-A recombinant protein and the biologically active fragment of AtPNP-A bind specifically to CAT2 in surface plasmon resonance (SPR) analyses, while a biologically inactive scrambled peptide does not. In vivo bimolecular fluorescence complementation (BiFC) showed that CAT2 interacts with AtPNP-A in chloroplasts. Furthermore, CAT2 activity is lower in homozygous atpnp-a knockdown compared with wild type plants, and atpnp-a knockdown plants phenocopy CAT2-deficient plants in their sensitivity to elevated H2O2, which is consistent with a direct modulatory effect of the PNP on the activity of CAT2 and hence H2O2 homeostasis. Our work underlines the critical role of AtPNP-A in modulating the activity of CAT2 and highlights a mechanism of fine-tuning plant responses to adverse conditions by PNPs.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
The plant natriuretic peptide receptor is a guanylyl cyclase and enables cGMP-dependent signaling.Plant Mol Biol. 2016 Jun;91(3):275-86. doi: 10.1007/s11103-016-0465-8. Epub 2016 Mar 5. Plant Mol Biol. 2016. PMID: 26945740
-
Plant natriuretic peptides are apoplastic and paracrine stress response molecules.Plant Cell Physiol. 2011 May;52(5):837-50. doi: 10.1093/pcp/pcr036. Epub 2011 Apr 7. Plant Cell Physiol. 2011. PMID: 21478192
-
The Arabidopsis thaliana natriuretic peptide AtPNP-A is a systemic regulator of leaf dark respiration and signals via the phloem.J Plant Physiol. 2011 Sep 15;168(14):1710-4. doi: 10.1016/j.jplph.2011.03.011. Epub 2011 May 7. J Plant Physiol. 2011. PMID: 21550130
-
Plant natriuretic peptides: systemic regulators of plant homeostasis and defense that can affect cardiomyoblasts.J Investig Med. 2013 Jun;61(5):823-6. doi: 10.2310/JIM.0b013e3182923395. J Investig Med. 2013. PMID: 23612149 Review.
-
Natriuretic peptides--a class of heterologous molecules in plants.Int J Biochem Cell Biol. 2003 Sep;35(9):1318-22. doi: 10.1016/s1357-2725(03)00032-3. Int J Biochem Cell Biol. 2003. PMID: 12798346 Review.
Cited by
-
Arabidopsis Plant Natriuretic Peptide Is a Novel Interactor of Rubisco Activase.Life (Basel). 2020 Dec 31;11(1):21. doi: 10.3390/life11010021. Life (Basel). 2020. PMID: 33396438 Free PMC article.
-
Amino acid motifs for the identification of novel protein interactants.Comput Struct Biotechnol J. 2022 Dec 10;21:326-334. doi: 10.1016/j.csbj.2022.12.012. eCollection 2023. Comput Struct Biotechnol J. 2022. PMID: 36582434 Free PMC article. Review.
-
Identification and Candidate Gene Evaluation of a Large Fast Neutron-Induced Deletion Associated with a High-Oil Phenotype in Soybean Seeds.Genes (Basel). 2024 Jul 8;15(7):892. doi: 10.3390/genes15070892. Genes (Basel). 2024. PMID: 39062671 Free PMC article.
-
Identification of a putative kinase interacting domain in the durum wheat catalase 1 (TdCAT1) protein.Heliyon. 2023 Aug 3;9(8):e18916. doi: 10.1016/j.heliyon.2023.e18916. eCollection 2023 Aug. Heliyon. 2023. PMID: 37609422 Free PMC article.
-
Computational Identification of Functional Centers in Complex Proteins: A Step-by-Step Guide With Examples.Front Bioinform. 2021 Mar 25;1:652286. doi: 10.3389/fbinf.2021.652286. eCollection 2021. Front Bioinform. 2021. PMID: 36303732 Free PMC article.
References
-
- Lemtiri-Chlieh, F., Thomas, L., Marondedze, C., Irving, H. & Gehring, C. Cyclic nucleotides and nucleotide cyclases in plant stress responses. In Abiotic Stress Response in Plants - Physiological, Biochemical and Genetic Perspectives (ed. Shanker, A.) 137–182 (InTech, 2012).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

