Genetic Spectrum of Syndromic and Non-Syndromic Hearing Loss in Pakistani Families
- PMID: 33187236
- PMCID: PMC7709052
- DOI: 10.3390/genes11111329
Genetic Spectrum of Syndromic and Non-Syndromic Hearing Loss in Pakistani Families
Abstract
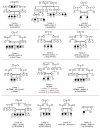
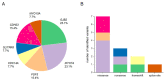
The current molecular genetic diagnostic rates for hereditary hearing loss (HL) vary considerably according to the population background. Pakistan and other countries with high rates of consanguineous marriages have served as a unique resource for studying rare and novel forms of recessive HL. A combined exome sequencing, bioinformatics analysis, and gene mapping approach for 21 consanguineous Pakistani families revealed 13 pathogenic or likely pathogenic variants in the genes GJB2, MYO7A, FGF3, CDC14A, SLITRK6, CDH23, and MYO15A, with an overall resolve rate of 61.9%. GJB2 and MYO7A were the most frequently involved genes in this cohort. All the identified variants were either homozygous or compound heterozygous, with two of them not previously described in the literature (15.4%). Overall, seven missense variants (53.8%), three nonsense variants (23.1%), two frameshift variants (15.4%), and one splice-site variant (7.7%) were observed. Syndromic HL was identified in five (23.8%) of the 21 families studied. This study reflects the extreme genetic heterogeneity observed in HL and expands the spectrum of variants in deafness-associated genes.
Keywords: Pakistan; consanguinity; exome sequencing; genetic diagnosis; genome-wide linkage analysis; hearing loss.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical

