Position-Dependent Differential Targeting of Somatic Hypermutation
- PMID: 33188076
- PMCID: PMC7726104
- DOI: 10.4049/jimmunol.2000496
Position-Dependent Differential Targeting of Somatic Hypermutation
Abstract
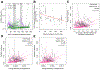
Somatic hypermutation (SHM) generates much of the Ab diversity necessary for affinity maturation and effective humoral immunity. The activation-induced cytidine deaminase-induced DNA lesions and error-prone repair that underlie SHM are known to exhibit intrinsic biases when targeting the Ig sequences. Computational models for SHM targeting often model the targeting probability of a nucleotide in a motif-based fashion, assuming that the same DNA motif is equally likely to be targeted regardless of its position along the Ig sequence. The validity of this assumption, however, has not been rigorously studied in vivo. In this study, by analyzing a large collection of 956,157 human Ig sequences while controlling for the confounding influence of selection, we show that the likelihood of a DNA 5-mer motif being targeted by SHM is not the same at different positions in the same Ig sequence. We found position-dependent differential SHM targeting for about three quarters of the 38 and 269 unique motifs from more than half of the 292 and 1912 motif-allele pairs analyzed using productive and nonproductive Ig sequences, respectively. The direction of the differential SHM targeting was largely conserved across individuals with no allele-specific effect within an IgH variable gene family, but was not consistent with general decay of SHM targeting with increasing distance from the transcription start site. However, SHM targeting did correlate positively with the mutability of the wider sequence neighborhood surrounding the motif. These findings provide insights and future directions for computational efforts toward modeling SHM.
Copyright © 2020 by The American Association of Immunologists, Inc.
Conflict of interest statement
Conflict of Interest Statement
S.H.K. receives consulting fees from Northrop Grumman.
Figures
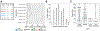



References
-
- Methot SP, and Di Noia JM. 2017. Molecular Mechanisms of Somatic Hypermutation and Class Switch Recombination. Adv. Immunol 133: 37–87. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

