Virulence-determinants and antibiotic-resistance genes of MDR-E. coli isolated from secondary infections following FMD-outbreak in cattle
- PMID: 33188216
- PMCID: PMC7666185
- DOI: 10.1038/s41598-020-75914-9
Virulence-determinants and antibiotic-resistance genes of MDR-E. coli isolated from secondary infections following FMD-outbreak in cattle
Abstract
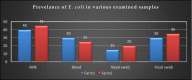
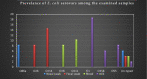
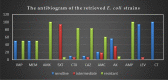
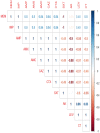
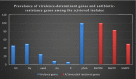
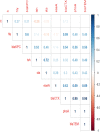
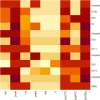
This study aimed to evaluate the prevalence, multidrug-resistance traits, PCR-detection of virulence, and antibiotic-resistance genes of E. coli isolated from secondary infections following FMD-outbreak in cattle. A total of 160 random samples were gathered from private dairy farms in Damietta Province, Egypt. The specimens were subjected to bacteriological examination, serotyping, congo-red binding assay, antibiogram-testing, and PCR-monitoring of virulence-determinant genes (tsh, phoA, hly, eaeA, sta, and lt) as well as the antibiotic-resistance genes (blaTEM, blaKPC, and blaCTX). The prevalence of E. coli was 30% (n = 48) distributed in 8 serogroups (40/48, 83.3%), while 8 isolates (8/48, 16.6%) were untypable. Besides, 83.3% of the examined isolates were positive for CR-binding. The tested strains harbored the virulence genes phoA, hly, tsh, eaeA, sta, and lt with a prevalence of 100% and 50%, 45.8%, 25%, 8.4%, and 6.2%, respectively. Furthermore, 50% of the recovered strains were multidrug-resistant (MDR) to penicillins, cephalosporins, and carbapenems, and are harboring the blaTEM, blaCTX, and blaKPC genes. Moreover, 25% of the examined strains are resistant to penicillins, and cephalosporins, and are harboring the blaTEM and blaCTX genes. To the best of our knowledge, this is the first report concerning the E. coli secondary bacterial infections following the FMD-outbreak. The emergence of MDR strains is considered a public health threat and indicates complicated treatment and bad prognosis of infections caused by such strains. Colistin sulfate and levofloxacin have a promising in vitro activity against MDR-E. coli.
Conflict of interest statement
The authors declare no competing interests.
Figures

References
-
- Verma AK, et al. Studies of the outbreaks of foot and mouth disease in Uttar Pradesh, India, between 2000 and 2006. Asian J. Epidemiol. 2011;3:141–147. doi: 10.3923/aje.2010.141.147. - DOI
-
- Verma AK, Kumar A, Rahal A, Bist B. Multi drug resistant pseudomonas aeruginosa: a secondary invader. Glob. J. Med. Res. 2014;14:6–9.
-
- Elbayoumy, M. K. et al.Molecular Characterization of Foot-and-Mouth Disease Virus Collected from Al-Fayoum and Beni-Suef Governorates in Egypt (2014).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials

