A GDSL lipase-like from Ipomoea batatas catalyzes efficient production of 3,5-diCQA when expressed in Pichia pastoris
- PMID: 33188250
- PMCID: PMC7666189
- DOI: 10.1038/s42003-020-01387-1
A GDSL lipase-like from Ipomoea batatas catalyzes efficient production of 3,5-diCQA when expressed in Pichia pastoris
Erratum in
-
Author Correction: A GDSL lipase-like from Ipomoea batatas catalyzes efficient production of 3,5-diCQA when expressed in Pichia pastoris.Commun Biol. 2020 Dec 2;3(1):746. doi: 10.1038/s42003-020-01488-x. Commun Biol. 2020. PMID: 33268810 Free PMC article.
Abstract
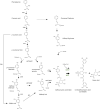
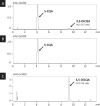
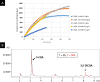
The synthesis of 3,5-dicaffeoylquinic acid (3,5-DiCQA) has attracted the interest of many researchers for more than 30 years. Recently, enzymes belonging to the BAHD acyltransferase family were shown to mediate its synthesis, albeit with notably low efficiency. In this study, a new enzyme belonging to the GDSL lipase-like family was identified and proven to be able to transform chlorogenic acid (5-O-caffeoylquinic acid, 5-CQA, CGA) in 3,5-DiCQA with a conversion rate of more than 60%. The enzyme has been produced in different expression systems but has only been shown to be active when transiently synthesized in Nicotiana benthamiana or stably expressed in Pichia pastoris. The synthesis of the molecule could be performed in vitro but also by a bioconversion approach beginning from pure 5-CQA or from green coffee bean extract, thereby paving the road for producing it on an industrial scale.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

