Gene and Protein Expression in Subjects With a Nystagmus-Associated AHR Mutation
- PMID: 33193710
- PMCID: PMC7542227
- DOI: 10.3389/fgene.2020.582796
Gene and Protein Expression in Subjects With a Nystagmus-Associated AHR Mutation
Abstract
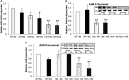
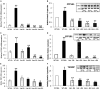
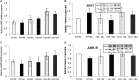
Recently, a consanguineous family was identified in Israel with three children affected by Infantile Nystagmus and Foveal Hypoplasia, following an autosomal recessive mode of inheritance. A homozygous stop mutation c.1861C > T; p.Q621∗ in the aryl hydrocarbon receptor (AHR) gene (AHR; MIM 600253) was identified that co-segregated with the disease in the larger family. AHR is the first gene to be identified causing an autosomal recessive Infantile Nystagmus-related disease in humans. The goal of this study is to delineate the molecular basis of this newly discovered human genetic disorder associated with a rare AHR gene mutation. The gene and protein expression levels of AHR and selected AHR targets from leukocyte cultures of healthy subjects and the patients were analyzed. We observed significant variation between mRNA and protein expression of CYP1A1, CYP1B1, and TiPARP under rest and AHR-induced conditions. The CYP1A1 enzymatic activity in induced leukocytes also differs significantly between the patients and healthy volunteers. Intriguingly, the heterozygous subjects demonstrate CYP1A1 and TiPARP gene and protein expression similar to homozygous patients. In contrast, CYP1B1 inducibility and expression vary between hetero- and homozygous subjects. Similarity and differences in gene and protein expression between heterozygotes and homozygous patients can give us a hint as to which metabolic pathway/s might be involved in the Nystagmus etiology. Thus, we have a unique human model for AHR deficiency that will allow us the opportunity to study the biochemical basis of this rare human mutation, as well as the involvement of AHR in other physiological processes.
Keywords: CYP1A1; CYP1B1; TiPARP; aryl hydrocarbon receptor; gene expression; human mutation; infantile nystagmus; protein expression.
Copyright © 2020 Borovok, Weiss, Sharkia, Reichenstein, Wissinger, Azem and Mahajnah.
Figures
Similar articles
-
Homozygous stop mutation in AHR causes autosomal recessive foveal hypoplasia and infantile nystagmus.Brain. 2019 Jun 1;142(6):1528-1534. doi: 10.1093/brain/awz098. Brain. 2019. PMID: 31009037 Free PMC article.
-
Correlation between gene expression of aryl hydrocarbon receptor (AhR), hydrocarbon receptor nuclear translocator (Arnt), cytochromes P4501A1 (CYP1A1) and 1B1 (CYP1B1), and inducibility of CYP1A1 and CYP1B1 in human lymphocytes.Toxicol Sci. 2003 Jan;71(1):20-6. doi: 10.1093/toxsci/71.1.20. Toxicol Sci. 2003. PMID: 12520072
-
Triclosan activates aryl hydrocarbon receptor (AhR)-dependent apoptosis and affects Cyp1a1 and Cyp1b1 expression in mouse neocortical neurons.Environ Res. 2016 Nov;151:106-114. doi: 10.1016/j.envres.2016.07.019. Epub 2016 Jul 27. Environ Res. 2016. PMID: 27474938
-
Constitutive regulation of CYP1B1 by the aryl hydrocarbon receptor (AhR) in pre-malignant and malignant mammary tissue.J Cell Biochem. 2008 May 15;104(2):402-17. doi: 10.1002/jcb.21630. J Cell Biochem. 2008. PMID: 18059014
-
Novel biallelic AHR splice site mutation cause isolated foveal hypoplasia in Saudi patient: a case report.Ophthalmic Genet. 2022 Jun;43(3):425-429. doi: 10.1080/13816810.2022.2039718. Epub 2022 Feb 21. Ophthalmic Genet. 2022. PMID: 35188035
Cited by
-
More than Meets the Eye: The Aryl Hydrocarbon Receptor is an Environmental Sensor, Physiological Regulator and a Therapeutic Target in Ocular Disease.Front Toxicol. 2022 Mar 3;4:791082. doi: 10.3389/ftox.2022.791082. eCollection 2022. Front Toxicol. 2022. PMID: 35295218 Free PMC article. Review.
References
-
- Atlas S. A., Vesell E. S., Nebert D. W. (1976). Genetic control of interindividual variations in the inducibility of aryl hydrocarbon hydroxylase in cultured human lymphocytes. Cancer Res. 36 4619–4630. - PubMed
LinkOut - more resources
Full Text Sources

