Hotspots of Human Mutation
- PMID: 33199048
- PMCID: PMC8366565
- DOI: 10.1016/j.tig.2020.10.003
Hotspots of Human Mutation
Abstract
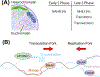
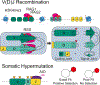
Mutation of the human genome results in three classes of genomic variation: single nucleotide variants; short insertions or deletions; and large structural variants (SVs). Some mutations occur during normal processes, such as meiotic recombination or B cell development, and others result from DNA replication or aberrant repair of breaks in sequence-specific contexts. Regardless of mechanism, mutations are subject to selection, and some hotspots can manifest in disease. Here, we discuss genomic regions prone to mutation, mechanisms contributing to mutation susceptibility, and the processes leading to their accumulation in normal and somatic genomes. With further, more accurate human genome sequencing, additional mutation hotspots, mechanistic details of their formation, and the relevance of hotspots to evolution and disease are likely to be discovered.
Keywords: DNA repair; indel; mutation hotspots; recurrent mutation; selection; structural variation.
Copyright © 2020 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures



References
-
- Gusella JF et al. (1983) A polymorphic DNA marker genetically linked to Huntington’s disease. Nature 306, 234–238 - PubMed
-
- Andrew SE et al. (1993) The relationship between trinucleotide (CAG) repeat length and clinical features of Huntington’s disease. Nat. Genet 4, 398–403 - PubMed
-
- MacDonald ME et al. (1993) A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

