Plant hairy roots enable high throughput identification of antimicrobials against Candidatus Liberibacter spp
- PMID: 33199718
- PMCID: PMC7669877
- DOI: 10.1038/s41467-020-19631-x
Plant hairy roots enable high throughput identification of antimicrobials against Candidatus Liberibacter spp
Abstract
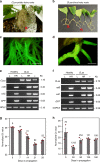
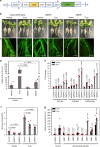
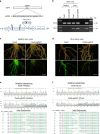
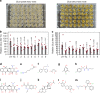
A major bottleneck in identifying therapies to control citrus greening and other devastating plant diseases caused by fastidious pathogens is our inability to culture the pathogens in defined media or axenic cultures. As such, conventional approaches for antimicrobial evaluation (genetic or chemical) rely on time-consuming, low-throughput and inherently variable whole-plant assays. Here, we report that plant hairy roots support the growth of fastidious pathogens like Candidatus Liberibacter spp., the presumptive causal agents of citrus greening, potato zebra chip and tomato vein greening diseases. Importantly, we leverage the microbial hairy roots for rapid, reproducible efficacy screening of multiple therapies. We identify six antimicrobial peptides, two plant immune regulators and eight chemicals which inhibit Candidatus Liberibacter spp. in plant tissues. The antimicrobials, either singly or in combination, can be used as near- and long-term therapies to control citrus greening, potato zebra chip and tomato vein greening diseases.
Conflict of interest statement
The Texas A&M University System has filed patent applications for the microbial hairy root system [15/353,645 and 16/54,178, status pending]. K.K.M., S.I., M.R., S.P., and P.N. are co-inventors on the related disclosures and patents. Southern Gardens Citrus (Clewiston, FL) has licensed rights in these patents and patent applications from The Texas A&M University System. A disclosure of invention for the compounds screened using this system and a U.S. provisional patent application (63031962) has been submitted. Southern Gardens Citrus also provided matching funds to the Foundation for Food and Agricultural Research New Innovator Award (2018-534,299) to K.K.M. Co-author M.S.I. (Director of Research and Business Development, Southern Gardens Citrus) contributed to the study design for screening antimicrobial peptides, data analysis, and preparation of the manuscript. The remaining authors declare no competing interests.
Figures

References
-
- Haapalainen M. Biology and epidemics of Candidatus Liberibacter species, psyllid-transmitted plant-pathogenic bacteria. Ann. Appl. Biol. 2014;165:172–198. doi: 10.1111/aab.12149. - DOI
-
- Munyaneza J. Zebra chip disease of potato: biology, epidemiology, and management. Am. Potato J. 2012;89:329–350. doi: 10.1007/s12230-012-9262-3. - DOI
-
- Greenway G. Economic impact of zebra chip control costs on grower returns in seven US states. Am. Potato J. 2014;91:714–719. doi: 10.1007/s12230-014-9404-x. - DOI
-
- CNAS. Economic impacts of zebra chip on the Texas potato industry. http://cnas.tamu.edu/zebra%20chip%20impacts%20final.pdf (Center for North American Studies, 2006).
-
- Bové JM. Huanglongbing: a destructive, newly-emerging, century-old disease of citrus. J. Plant Pathol. 2006;88:7–37.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

