The conformational cycle of a prototypical voltage-gated sodium channel
- PMID: 33199904
- PMCID: PMC7678813
- DOI: 10.1038/s41589-020-0644-4
The conformational cycle of a prototypical voltage-gated sodium channel
Abstract
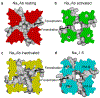
Electrical signaling was a dramatic development in evolution, allowing complex single-cell organisms like Paramecium to coordinate movement and early metazoans like worms and jellyfish to send regulatory signals rapidly over increasing distances. But how are electrical signals generated in biology? In fact, voltage-gated sodium channels conduct sodium currents that initiate electrical signals in all kingdoms of life, from bacteria to man. They are responsible for generating the action potential in vertebrate nerve and muscle, neuroendocrine cells, and other cell types1,2. Because of the high level of conservation of their core structure, it is likely that their fundamental mechanisms of action are conserved as well. Here we describe the complete cycle of conformational changes that a bacterial sodium channel undergoes as it transitions from resting to activated/open and inactivated/closed states, based on high-resolution structural studies of a single sodium channel. We further relate this conformational cycle of the ancestral sodium channel to the function of its vertebrate orthologs. The strong conservation of amino acid sequence and three-dimensional structure suggests that this model, at a fundamental level, is relevant for both prokaryotic and eukaryotic sodium channels, as well as voltage-gated calcium and potassium channels.
Conflict of interest statement
Competing interests
The authors declare no competing interests.
Figures
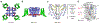



Similar articles
-
Biophysical Adaptations of Prokaryotic Voltage-Gated Sodium Channels.Curr Top Membr. 2016;78:39-64. doi: 10.1016/bs.ctm.2015.12.003. Epub 2016 Feb 1. Curr Top Membr. 2016. PMID: 27586280
-
Structure-based assessment of disease-related mutations in human voltage-gated sodium channels.Protein Cell. 2017 Jun;8(6):401-438. doi: 10.1007/s13238-017-0372-z. Epub 2017 Feb 1. Protein Cell. 2017. PMID: 28150151 Free PMC article. Review.
-
A native prokaryotic voltage-dependent calcium channel with a novel selectivity filter sequence.Elife. 2020 Feb 25;9:e52828. doi: 10.7554/eLife.52828. Elife. 2020. PMID: 32093827 Free PMC article.
-
Voltage-gated sodium channels assemble and gate as dimers.Nat Commun. 2017 Dec 12;8(1):2077. doi: 10.1038/s41467-017-02262-0. Nat Commun. 2017. PMID: 29233994 Free PMC article.
-
Molecular evolution of K+ channels in primitive eukaryotes.Soc Gen Physiol Ser. 1994;49:213-22. Soc Gen Physiol Ser. 1994. PMID: 7939896 Review.
Cited by
-
Functional Characterization of Four Known Cav2.1 Variants Associated with Neurodevelopmental Disorders.Membranes (Basel). 2023 Jan 11;13(1):96. doi: 10.3390/membranes13010096. Membranes (Basel). 2023. PMID: 36676903 Free PMC article.
-
The differential impacts of equivalent gating-charge mutations in voltage-gated sodium channels.J Gen Physiol. 2025 Mar 3;157(2):e202413669. doi: 10.1085/jgp.202413669. Epub 2025 Jan 17. J Gen Physiol. 2025. PMID: 39820972
-
Harnessing AlphaFold to reveal hERG channel conformational state secrets.Elife. 2025 Jul 14;13:RP104901. doi: 10.7554/eLife.104901. Elife. 2025. PMID: 40658102 Free PMC article.
-
Characterizing fenestration size in sodium channel subtypes and their accessibility to inhibitors.Biophys J. 2022 Jan 18;121(2):193-206. doi: 10.1016/j.bpj.2021.12.025. Epub 2021 Dec 24. Biophys J. 2022. PMID: 34958776 Free PMC article.
-
Deciphering Ca2+ permeation and valence selectivity in CaV1: Molecular dynamics simulations reveal the three-ion knock-on mechanism.Proc Natl Acad Sci U S A. 2025 Jun 3;122(22):e2424694122. doi: 10.1073/pnas.2424694122. Epub 2025 May 29. Proc Natl Acad Sci U S A. 2025. PMID: 40440072 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

