Inhibition of the de novo pyrimidine biosynthesis pathway limits ribosomal RNA transcription causing nucleolar stress in glioblastoma cells
- PMID: 33201894
- PMCID: PMC7707548
- DOI: 10.1371/journal.pgen.1009117
Inhibition of the de novo pyrimidine biosynthesis pathway limits ribosomal RNA transcription causing nucleolar stress in glioblastoma cells
Abstract
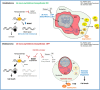
Glioblastoma is the most common and aggressive type of cancer in the brain; its poor prognosis is often marked by reoccurrence due to resistance to the chemotherapeutic agent temozolomide, which is triggered by an increase in the expression of DNA repair enzymes such as MGMT. The poor prognosis and limited therapeutic options led to studies targeted at understanding specific vulnerabilities of glioblastoma cells. Metabolic adaptations leading to increased synthesis of nucleotides by de novo biosynthesis pathways are emerging as key alterations driving glioblastoma growth. In this study, we show that enzymes necessary for the de novo biosynthesis of pyrimidines, DHODH and UMPS, are elevated in high grade gliomas and in glioblastoma cell lines. We demonstrate that DHODH's activity is necessary to maintain ribosomal DNA transcription (rDNA). Pharmacological inhibition of DHODH with the specific inhibitors brequinar or ML390 effectively depleted the pool of pyrimidines in glioblastoma cells grown in vitro and in vivo and impaired rDNA transcription, leading to nucleolar stress. Nucleolar stress was visualized by the aberrant redistribution of the transcription factor UBF and the nucleolar organizer nucleophosmin 1 (NPM1), as well as the stabilization of the transcription factor p53. Moreover, DHODH inhibition decreased the proliferation of glioblastoma cells, including temozolomide-resistant cells. Importantly, the addition of exogenous uridine, which reconstitutes the cellular pool of pyrimidine by the salvage pathway, to the culture media recovered the impaired rDNA transcription, nucleolar morphology, p53 levels, and proliferation of glioblastoma cells caused by the DHODH inhibitors. Our in vivo data indicate that while inhibition of DHODH caused a dramatic reduction in pyrimidines in tumor cells, it did not affect the overall pyrimidine levels in normal brain and liver tissues, suggesting that pyrimidine production by the salvage pathway may play an important role in maintaining these nucleotides in normal cells. Our study demonstrates that glioblastoma cells heavily rely on the de novo pyrimidine biosynthesis pathway to generate ribosomal RNA (rRNA) and thus, we identified an approach to inhibit ribosome production and consequently the proliferation of glioblastoma cells through the specific inhibition of the de novo pyrimidine biosynthesis pathway.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

