Loss of TAX1BP1-Directed Autophagy Results in Protein Aggregate Accumulation in the Brain
- PMID: 33207181
- PMCID: PMC7771836
- DOI: 10.1016/j.molcel.2020.10.041
Loss of TAX1BP1-Directed Autophagy Results in Protein Aggregate Accumulation in the Brain
Erratum in
-
Loss of TAX1BP1-Directed Autophagy Results in Protein Aggregate Accumulation in the Brain.Mol Cell. 2022 Apr 7;82(7):1383-1385. doi: 10.1016/j.molcel.2022.03.020. Mol Cell. 2022. PMID: 35395199 Free PMC article. No abstract available.
Abstract
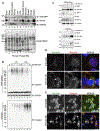
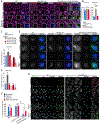
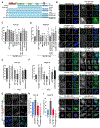
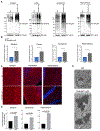
Protein aggregates disrupt cellular homeostasis, causing toxicity linked to neurodegeneration. Selective autophagic elimination of aggregates is critical to protein quality control, but how aggregates are selectively targeted for degradation is unclear. We compared the requirements for autophagy receptor proteins: OPTN, NBR1, p62, NDP52, and TAX1BP1 in clearance of proteotoxic aggregates. Endogenous TAX1BP1 is recruited to and required for the clearance of stress-induced aggregates, whereas ectopic expression of TAX1BP1 increases clearance through autophagy, promoting viability of human induced pluripotent stem cell-derived neurons. In contrast, TAX1BP1 depletion sensitizes cells to several forms of aggregate-induced proteotoxicity. Furthermore, TAX1BP1 is more specifically expressed in the brain compared to other autophagy receptor proteins. In vivo, loss of TAX1BP1 results in accumulation of high molecular weight ubiquitin conjugates and premature lipofuscin accumulation in brains of young TAX1BP1 knockout mice. TAX1BP1 mediates clearance of a broad range of cytotoxic proteins indicating therapeutic potential in neurodegenerative diseases.
Keywords: Huntington’s disease; OPTN; aggregate; aggrephagy; autophagy receptor; p62; proteostasis; proteotoxic stress; selective autophagy; ubiquitin.
Published by Elsevier Inc.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures



Similar articles
-
CCDC50 mediates the clearance of protein aggregates to prevent cellular proteotoxicity.Autophagy. 2024 Nov;20(11):2529-2539. doi: 10.1080/15548627.2024.2367183. Epub 2024 Jul 30. Autophagy. 2024. PMID: 38869076 Free PMC article.
-
TAX1BP1 and FIP200 orchestrate non-canonical autophagy of p62 aggregates for mouse neural stem cell maintenance.Zool Res. 2024 Jul 18;45(4):937-950. doi: 10.24272/j.issn.2095-8137.2024.021. Zool Res. 2024. PMID: 39021082 Free PMC article.
-
CCT2 is an aggrephagy receptor for clearance of solid protein aggregates.Cell. 2022 Apr 14;185(8):1325-1345.e22. doi: 10.1016/j.cell.2022.03.005. Epub 2022 Apr 1. Cell. 2022. PMID: 35366418
-
Advances in Aggrephagy: Mechanisms, Disease Implications, and Therapeutic Strategies.J Cell Physiol. 2025 Jan;240(1):e31512. doi: 10.1002/jcp.31512. J Cell Physiol. 2025. PMID: 39749851 Review.
-
Multifaceted roles of TAX1BP1 in autophagy.Autophagy. 2023 Jan;19(1):44-53. doi: 10.1080/15548627.2022.2070331. Epub 2022 May 9. Autophagy. 2023. PMID: 35470757 Free PMC article. Review.
Cited by
-
Modeling native and seeded Synuclein aggregation and related cellular dysfunctions in dopaminergic neurons derived by a new set of isogenic iPSC lines with SNCA multiplications.Cell Death Dis. 2022 Oct 19;13(10):881. doi: 10.1038/s41419-022-05330-6. Cell Death Dis. 2022. PMID: 36261424 Free PMC article.
-
An RB1CC1 Missense Variant in Nova Scotia Duck Tolling Retrievers with Degenerative Encephalopathy.Genes (Basel). 2025 Feb 25;16(3):269. doi: 10.3390/genes16030269. Genes (Basel). 2025. PMID: 40149422 Free PMC article.
-
NDP52 and its emerging role in pathogenesis.Cell Death Dis. 2025 May 3;16(1):359. doi: 10.1038/s41419-025-07668-z. Cell Death Dis. 2025. PMID: 40319017 Free PMC article. Review.
-
p62 and NBR1 functions are dispensable for aggrephagy in mouse ESCs and ESC-derived neurons.Life Sci Alliance. 2023 Aug 24;6(11):e202301936. doi: 10.26508/lsa.202301936. Print 2023 Nov. Life Sci Alliance. 2023. PMID: 37620146 Free PMC article.
-
TNIP1 inhibits selective autophagy via bipartite interaction with LC3/GABARAP and TAX1BP1.Mol Cell. 2023 Mar 16;83(6):927-941.e8. doi: 10.1016/j.molcel.2023.02.023. Epub 2023 Mar 9. Mol Cell. 2023. PMID: 36898370 Free PMC article.
References
-
- Balchin D, Hayer-Hartl M, and Hartl FU (2016). In vivo aspects of protein folding and quality control. Science 353, aac4354. - PubMed
-
- Bennett EJ, Bence NF, Jayakumar R, and Kopito RR (2005). Global impairment of the ubiquitin-proteasome system by nuclear or cytoplasmic protein aggregates precedes inclusion body formation. Mol. Cell 17,351–365. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

