Whole Genome Sequencing and Tn 5-Insertion Mutagenesis of Pseudomonas taiwanensis CMS to Probe Its Antagonistic Activity Against Rice Bacterial Blight Disease
- PMID: 33207795
- PMCID: PMC7696974
- DOI: 10.3390/ijms21228639
Whole Genome Sequencing and Tn 5-Insertion Mutagenesis of Pseudomonas taiwanensis CMS to Probe Its Antagonistic Activity Against Rice Bacterial Blight Disease
Abstract
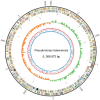
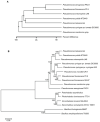
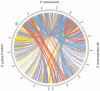
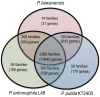
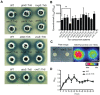
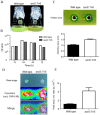
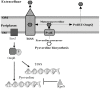
The Gram-negative bacterium Pseudomonas taiwanensis is a novel bacterium that uses shrimp shell waste as its sole sources of carbon and nitrogen. It is a versatile bacterium with potential for use in biological control, with activities including toxicity toward insects, fungi, and the rice pathogen Xanthomonas oryzae pv.oryzae (Xoo). In this study, the complete 5.08-Mb genome sequence of P. taiwanensis CMS was determined by a combination of NGS/Sanger sequencing and optical mapping. Comparison of optical maps of seven Pseudomonas species showed that P. taiwanensis is most closely related to P. putida KT 2400. We screened a total of 11,646 individual Tn5-transponson tagged strains to identify genes that are involved in the production and regulation of the iron-chelator pyoverdine in P. taiwanensis, which is a key anti-Xoo factor. Our results indicated that the two-component system (TCS) EnvZ/OmpR plays a positive regulatory role in the production of pyoverdine, whereas the sigma factor RpoS functions as a repressor. The knowledge of the molecular basis of the regulation of pyoverdine by P. taiwanensis provided herein will be useful for its development for use in biological control, including as an anti-Xoo agent.
Keywords: Pseudomonas taiwanensis; Rice bacterial blight; Xanthomonas oryzae pv.oryzae (Xoo); biocontrol agent.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Involvement of type VI secretion system in secretion of iron chelator pyoverdine in Pseudomonas taiwanensis.Sci Rep. 2016 Sep 8;6:32950. doi: 10.1038/srep32950. Sci Rep. 2016. PMID: 27605490 Free PMC article.
-
EcpA, an extracellular protease, is a specific virulence factor required by Xanthomonas oryzae pv. oryzicola but not by X. oryzae pv. oryzae in rice.Microbiology (Reading). 2012 Sep;158(Pt 9):2372-2383. doi: 10.1099/mic.0.059964-0. Epub 2012 Jun 14. Microbiology (Reading). 2012. PMID: 22700650
-
CitB is required for full virulence of Xanthomonas oryzae pv. oryzae.World J Microbiol Biotechnol. 2015 Oct;31(10):1619-27. doi: 10.1007/s11274-015-1914-2. Epub 2015 Aug 7. World J Microbiol Biotechnol. 2015. PMID: 26250545
-
Isolation and characterization of a novel phage Xoo-sp2 that infects Xanthomonas oryzae pv. oryzae.J Gen Virol. 2018 Oct;99(10):1453-1462. doi: 10.1099/jgv.0.001133. Epub 2018 Aug 10. J Gen Virol. 2018. PMID: 30102145
-
[Identification of a hrp-associated gene hpaB in the role of pathogenicity of Xanthomonas oryzae pv. oryzae].Wei Sheng Wu Xue Bao. 2007 Jun;47(3):402-6. Wei Sheng Wu Xue Bao. 2007. PMID: 17672295 Chinese.
Cited by
-
Disease-resistant varieties of Chinese cabbage (Brassica rapa L. ssp. pekinensis) inhibit Plasmodiophora brassicae infestation by stabilising root flora structure.Front Plant Sci. 2024 Mar 5;15:1328845. doi: 10.3389/fpls.2024.1328845. eCollection 2024. Front Plant Sci. 2024. PMID: 38504895 Free PMC article.
References
-
- Buell C.R., Joardar V., Lindeberg M., Selengut J., Paulsen I.T., Gwinn M.L., Dodson R.J., Deboy R.T., Durkin A.S., Kolonay J.F., et al. The complete genome sequence of the Arabidopsis and tomato pathogen Pseudomonas syringae pv. tomato DC3000. Proc. Natl. Acad. Sci. USA. 2003;100:10181–10186. doi: 10.1073/pnas.1731982100. - DOI - PMC - PubMed
-
- Vodovar N., Vallenet D., Cruveiller S., Rouy Z., Barbe V., Acosta C., Cattolico L., Jubin C., Lajus A., Segurens B., et al. Complete genome sequence of the entomopathogenic and metabolically versatile soil bacterium Pseudomonas entomophila. Nat. Biotechnol. 2006;24:673–679. doi: 10.1038/nbt1212. - DOI - PubMed
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Miscellaneous

