Long-term skin-resident memory T cells proliferate in situ and are involved in human graft-versus-host disease
- PMID: 33208504
- PMCID: PMC7615006
- DOI: 10.1126/scitranslmed.abb7028
Long-term skin-resident memory T cells proliferate in situ and are involved in human graft-versus-host disease
Abstract
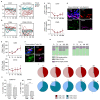
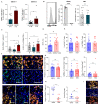
The skin contains a population of tissue-resident memory T cells (Trm) that is thought to contribute to local tissue homeostasis and protection against environmental injuries. Although information about the regulation, survival program, and pathophysiological roles of Trm has been obtained from murine studies, little is known about the biology of human cutaneous Trm Here, we showed that host-derived CD69+ αβ memory T cell clones in the epidermis and dermis remain stable and functionally competent for at least 10 years in patients with allogeneic hematopoietic stem cell transplantation. Single-cell RNA sequencing revealed low expression of genes encoding tissue egress molecules by long-term persisting Trm in the skin, whereas tissue retention molecules and stem cell markers were displayed by Trm The transcription factor RUNX3 and the surface molecule galectin-3 were preferentially expressed by host T cells at the RNA and protein levels, suggesting two new markers for human skin Trm Furthermore, skin lesions from patients developing graft-versus-host disease (GVHD) showed a large number of cytokine-producing host-derived Trm, suggesting a contribution of these cells to the pathogenesis of GVHD. Together, our studies highlighted the relationship between the local human skin environment and long-term persisting Trm, which differs from murine skin. Our results also indicated that local tissue inflammation occurs through host-derived Trm after allogeneic hematopoietic stem cell transplantation.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Figures





References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

