Structural and dynamical insights into the PH domain of p62 in human TFIIH
- PMID: 33211877
- PMCID: PMC7969019
- DOI: 10.1093/nar/gkaa1045
Structural and dynamical insights into the PH domain of p62 in human TFIIH
Abstract
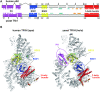
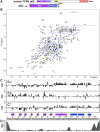
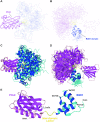
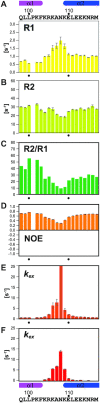
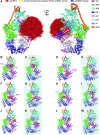
TFIIH is a crucial transcription and DNA repair factor consisting of the seven-subunit core. The core subunit p62 contains a pleckstrin homology domain (PH-D), which is essential for locating TFIIH at transcription initiation and DNA damage sites, and two BSD (BTF2-like transcription factors, synapse-associated proteins and DOS2-like proteins) domains. A recent cryo-electron microscopy (cryo-EM) structure of human TFIIH visualized most parts of core, except for the PH-D. Here, by nuclear magnetic resonance spectroscopy we have established the solution structure of human p62 PH-D connected to the BSD1 domain by a highly flexible linker, suggesting the flexibility of PH-D in TFIIH. Based on this dynamic character, the PH-D was modeled in the cryo-EM structure to obtain the whole human TFIIH core structure, which indicates that the PH-D moves around the surface of core with a specific but limited spatial distribution; these dynamic structures were refined by molecular dynamics (MD) simulations. Furthermore, we built models, also refined by MD simulations, of TFIIH in complex with five p62-binding partners, including transcription factors TFIIEα, p53 and DP1, and nucleotide excision repair factors XPC and UVSSA. The models explain why the PH-D is crucially targeted by these factors, which use their intrinsically disordered acidic regions for TFIIH recruitment.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
Dynamic structures of intrinsically disordered proteins related to the general transcription factor TFIIH, nucleosomes, and histone chaperones.Biophys Rev. 2022 Nov 18;14(6):1449-1472. doi: 10.1007/s12551-022-01014-9. eCollection 2022 Dec. Biophys Rev. 2022. PMID: 36659983 Free PMC article. Review.
-
Common TFIIH recruitment mechanism in global genome and transcription-coupled repair subpathways.Nucleic Acids Res. 2017 Dec 15;45(22):13043-13055. doi: 10.1093/nar/gkx970. Nucleic Acids Res. 2017. PMID: 29069470 Free PMC article.
-
Structural polymorphism of the PH domain in TFIIH.Biosci Rep. 2023 Jul 26;43(7):BSR20230846. doi: 10.1042/BSR20230846. Biosci Rep. 2023. PMID: 37340985 Free PMC article.
-
Structural insight into the TFIIE-TFIIH interaction: TFIIE and p53 share the binding region on TFIIH.EMBO J. 2008 Apr 9;27(7):1161-71. doi: 10.1038/emboj.2008.47. Epub 2008 Mar 20. EMBO J. 2008. PMID: 18354501 Free PMC article.
-
High-resolution cryo-EM structures of TFIIH and their functional implications.Curr Opin Struct Biol. 2019 Dec;59:188-194. doi: 10.1016/j.sbi.2019.08.002. Epub 2019 Oct 7. Curr Opin Struct Biol. 2019. PMID: 31600675 Free PMC article. Review.
Cited by
-
Targeting protein-protein interactions in the DNA damage response pathways for cancer chemotherapy.RSC Chem Biol. 2021 Jun 21;2(4):1167-1195. doi: 10.1039/d1cb00101a. eCollection 2021 Aug 5. RSC Chem Biol. 2021. PMID: 34458830 Free PMC article. Review.
-
Dynamic structures of intrinsically disordered proteins related to the general transcription factor TFIIH, nucleosomes, and histone chaperones.Biophys Rev. 2022 Nov 18;14(6):1449-1472. doi: 10.1007/s12551-022-01014-9. eCollection 2022 Dec. Biophys Rev. 2022. PMID: 36659983 Free PMC article. Review.
-
Modulation of the Nrf-2 and HO-1 signalling axis is associated with Betaine's abatement of fluoride-induced hepatorenal toxicities in rats.Naunyn Schmiedebergs Arch Pharmacol. 2024 Oct;397(10):7725-7745. doi: 10.1007/s00210-024-03133-4. Epub 2024 May 7. Naunyn Schmiedebergs Arch Pharmacol. 2024. PMID: 38713257
-
Three human RNA polymerases interact with TFIIH via a common RPB6 subunit.Nucleic Acids Res. 2022 Jan 11;50(1):1-16. doi: 10.1093/nar/gkab612. Nucleic Acids Res. 2022. PMID: 34268577 Free PMC article.
-
Integrative approaches for characterizing protein dynamics: NMR, CryoEM, and computer simulations.Curr Opin Struct Biol. 2024 Feb;84:102736. doi: 10.1016/j.sbi.2023.102736. Epub 2023 Dec 3. Curr Opin Struct Biol. 2024. PMID: 38048753 Free PMC article. Review.
References
-
- Feaver W.J., Gileadi O., Kornberg R.D.. Purification and characterization of yeast RNA polymerase II transcription factor b. J. Biol. Chem. 1991; 266:19000–19005. - PubMed
-
- Gerard M., Fischer L., Moncollin V., Chipoulet J.M., Chambon P., Egly J.M.. Purification and interaction properties of the human RNA polymerase B(II) general transcription factor BTF2. J. Biol. Chem. 1991; 266:20940–20945. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

