A Novel Purification Procedure for Active Recombinant Human DPP4 and the Inability of DPP4 to Bind SARS-CoV-2
- PMID: 33218025
- PMCID: PMC7698748
- DOI: 10.3390/molecules25225392
A Novel Purification Procedure for Active Recombinant Human DPP4 and the Inability of DPP4 to Bind SARS-CoV-2
Abstract
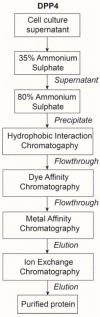
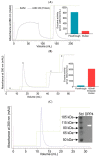
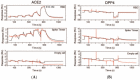
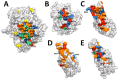
Proteases catalyse irreversible posttranslational modifications that often alter a biological function of the substrate. The protease dipeptidyl peptidase 4 (DPP4) is a pharmacological target in type 2 diabetes therapy primarily because it inactivates glucagon-like protein-1. DPP4 also has roles in steatosis, insulin resistance, cancers and inflammatory and fibrotic diseases. In addition, DPP4 binds to the spike protein of the MERS virus, causing it to be the human cell surface receptor for that virus. DPP4 has been identified as a potential binding target of SARS-CoV-2 spike protein, so this question requires experimental investigation. Understanding protein structure and function requires reliable protocols for production and purification. We developed such strategies for baculovirus generated soluble recombinant human DPP4 (residues 29-766) produced in insect cells. Purification used differential ammonium sulphate precipitation, hydrophobic interaction chromatography, dye affinity chromatography in series with immobilised metal affinity chromatography, and ion-exchange chromatography. The binding affinities of DPP4 to the SARS-CoV-2 full-length spike protein and its receptor-binding domain (RBD) were measured using surface plasmon resonance and ELISA. This optimised DPP4 purification procedure yielded 1 to 1.8 mg of pure fully active soluble DPP4 protein per litre of insect cell culture with specific activity >30 U/mg, indicative of high purity. No specific binding between DPP4 and CoV-2 spike protein was detected by surface plasmon resonance or ELISA. In summary, a procedure for high purity high yield soluble human DPP4 was achieved and used to show that, unlike MERS, SARS-CoV-2 does not bind human DPP4.
Keywords: Covid-19; DPP4; protease; recombinant protein.
Conflict of interest statement
The authors declare no conflict of interest. DPP4 enzyme produced as described here is sold by Sigma-Aldrich, with catalogue number D4943, with a financial benefit to the research led by M.D.G.
Figures

Similar articles
-
Recombinant Receptor-Binding Domains of Multiple Middle East Respiratory Syndrome Coronaviruses (MERS-CoVs) Induce Cross-Neutralizing Antibodies against Divergent Human and Camel MERS-CoVs and Antibody Escape Mutants.J Virol. 2016 Dec 16;91(1):e01651-16. doi: 10.1128/JVI.01651-16. Print 2017 Jan 1. J Virol. 2016. PMID: 27795425 Free PMC article.
-
Does the SARS-CoV-2 Spike Protein Receptor Binding Domain Interact Effectively with the DPP4 (CD26) Receptor? A Molecular Docking Study.Int J Mol Sci. 2021 Jun 29;22(13):7001. doi: 10.3390/ijms22137001. Int J Mol Sci. 2021. PMID: 34209788 Free PMC article.
-
Adaptive Evolution of MERS-CoV to Species Variation in DPP4.Cell Rep. 2018 Aug 14;24(7):1730-1737. doi: 10.1016/j.celrep.2018.07.045. Cell Rep. 2018. PMID: 30110630 Free PMC article.
-
DPP4 and ACE2 in Diabetes and COVID-19: Therapeutic Targets for Cardiovascular Complications?Front Pharmacol. 2020 Aug 7;11:1161. doi: 10.3389/fphar.2020.01161. eCollection 2020. Front Pharmacol. 2020. PMID: 32848769 Free PMC article. Review.
-
Revolutionizing Treatment Strategies for Autoimmune and Inflammatory Disorders: The Impact of Dipeptidyl-Peptidase 4 Inhibitors.J Inflamm Res. 2024 Mar 23;17:1897-1917. doi: 10.2147/JIR.S442106. eCollection 2024. J Inflamm Res. 2024. PMID: 38544813 Free PMC article. Review.
Cited by
-
Proline-specific peptidase activities (DPP4, PRCP, FAP and PREP) in plasma of hospitalized COVID-19 patients.Clin Chim Acta. 2022 Jun 1;531:4-11. doi: 10.1016/j.cca.2022.03.005. Epub 2022 Mar 10. Clin Chim Acta. 2022. PMID: 35283094 Free PMC article.
-
ACE2 function in the pancreatic islet: Implications for relationship between SARS-CoV-2 and diabetes.Acta Physiol (Oxf). 2021 Dec;233(4):e13733. doi: 10.1111/apha.13733. Epub 2021 Oct 2. Acta Physiol (Oxf). 2021. PMID: 34561952 Free PMC article. Review.
-
ACE2-Fc and DPP4-Fc decoy receptors against SARS-CoV-2 and MERS-CoV variants: a quick therapeutic option for current and future coronaviruses outbreaks.Antib Ther. 2023 Dec 12;7(1):53-66. doi: 10.1093/abt/tbad030. eCollection 2024 Jan. Antib Ther. 2023. PMID: 38371953 Free PMC article.
-
Identification of Transcription Factors Regulating SARS-CoV-2 Tropism Factor Expression by Inferring Cell-Type-Specific Transcriptional Regulatory Networks in Human Lungs.Viruses. 2022 Apr 17;14(4):837. doi: 10.3390/v14040837. Viruses. 2022. PMID: 35458567 Free PMC article.
-
Increased complications of COVID-19 in people with cardiovascular disease: Role of the renin-angiotensin-aldosterone system (RAAS) dysregulation.Chem Biol Interact. 2022 Jan 5;351:109738. doi: 10.1016/j.cbi.2021.109738. Epub 2021 Nov 3. Chem Biol Interact. 2022. PMID: 34740598 Free PMC article. Review.
References
-
- Rosenstock J., Perkovic V., Johansen O.E., Cooper M.E., Kahn S.E., Marx N., McGuire D.K. Effect of Linagliptin vs. Placebo on Major Cardiovascular Events in Adults with Type 2 Diabetes and High Cardiovascular and Renal Risk: The CARMELINA Randomized Clinical Trial. JAMA. 2019;321:69–79. doi: 10.1001/jama.2018.18269. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

