Nanoparticle-complexed antimiRs for inhibiting tumor growth and metastasis in prostate carcinoma and melanoma
- PMID: 33228711
- PMCID: PMC7685669
- DOI: 10.1186/s12951-020-00728-w
Nanoparticle-complexed antimiRs for inhibiting tumor growth and metastasis in prostate carcinoma and melanoma
Abstract
Background: MiRNAs act as negative regulators of gene expression through target mRNA degradation or inhibition of its translation. In cancer, several miRNAs are upregulated and play crucial roles in tumorigenesis, making the inhibition of these oncomiRs an interesting therapeutic approach. This can be achieved by directly complementary single-stranded anti-miRNA oligonucleotides (antimiRs). A major bottleneck in antimiR therapy, however, is their efficient delivery. The nanoparticle formation with polyethylenimine (PEI) may be particularly promising, based on the PEI's ability to electrostatically interact with oligonucleotides. This leads to their protection and supports delivery. In the present study, we explore for the first time PEI for antimiR formulation and delivery. We use the branched low molecular weight PEI F25-LMW for the complexation of different antimiRs, and analyse tumor- and metastasis-inhibitory effects of PEI/antimiR complexes in different tumor models.
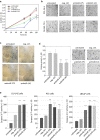
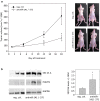
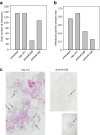
Results: In prostate carcinoma, transfection of antimiRs against miR-375 and miR-141 leads to tumor cell inhibition in 2D- and 3D-models. More importantly, an in vivo tumor therapy study in prostate carcinoma xenografts reveals anti-tumor effects of the PEI/antimiR complexes. In advanced melanoma and metastasis, we identify by a microRNA screen miR-150 as a particularly relevant oncomiR candidate, and validate this result in vitro and in vivo. Again, the systemic application of PEI/antimiR complexes inhibiting this miRNA, or the previously described antimiR-638, leads to profound tumor growth inhibition. These effects are associated with the upregulation of direct miRNA target genes. In a melanoma metastasis mouse model, anti-metastatic effects of PEI/antimiR treatment are observed as well.
Conclusions: We thus describe PEI-based complexes as efficient platform for antimiR therapy, as determined in two different tumor entities using in vivo models of tumor growth or metastasis. Our study also highlights the therapeutic relevance of miR-375, miR-141, miR-150 and miR-638 as target miRNAs for antimiR-mediated inhibition.
Keywords: Antimir; PEI; PEI/antimiR nanoparticles; Polyethylenimine; Therapeutic miRNA inhibition.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



Similar articles
-
Extracellular vesicle (ECV)-modified polyethylenimine (PEI) complexes for enhanced siRNA delivery in vitro and in vivo.J Control Release. 2020 Mar 10;319:63-76. doi: 10.1016/j.jconrel.2019.12.032. Epub 2019 Dec 19. J Control Release. 2020. PMID: 31866504
-
PEI-complexed LNA antiseeds as miRNA inhibitors.RNA Biol. 2012 Aug;9(8):1088-98. doi: 10.4161/rna.21165. Epub 2012 Aug 1. RNA Biol. 2012. PMID: 22894918 Free PMC article.
-
MicroRNA replacement therapy for miR-145 and miR-33a is efficacious in a model of colon carcinoma.Cancer Res. 2011 Aug 1;71(15):5214-24. doi: 10.1158/0008-5472.CAN-10-4645. Epub 2011 Jun 20. Cancer Res. 2011. PMID: 21690566
-
Bioresponsive Nanoparticles Boost Starvation Therapy and Prevent Premetastatic Niche Formation for Pulmonary Metastasis Treatment.ACS Appl Mater Interfaces. 2024 Oct 2;16(39):51798-51806. doi: 10.1021/acsami.4c11686. Epub 2024 Sep 20. ACS Appl Mater Interfaces. 2024. PMID: 39301793 Review.
-
Canonical and non-canonical barriers facing antimiR cancer therapeutics.Curr Med Chem. 2013;20(29):3582-93. doi: 10.2174/0929867311320290004. Curr Med Chem. 2013. PMID: 23745563 Free PMC article. Review.
Cited by
-
Integrated Analysis of Mutations, miRNA and mRNA Expression in Glioblastoma.Int J Gen Med. 2021 Nov 16;14:8281-8292. doi: 10.2147/IJGM.S336421. eCollection 2021. Int J Gen Med. 2021. PMID: 34815700 Free PMC article.
-
Identification of MicroRNAs as Viable Aggressiveness Biomarkers for Prostate Cancer.Biomedicines. 2021 Jun 5;9(6):646. doi: 10.3390/biomedicines9060646. Biomedicines. 2021. PMID: 34198846 Free PMC article.
-
Circulating MicroRNAs: functional biomarkers for melanoma prognosis and treatment.Mol Cancer. 2025 Mar 28;24(1):99. doi: 10.1186/s12943-025-02298-7. Mol Cancer. 2025. PMID: 40156012 Free PMC article. Review.
-
Current Landscape and Future Directions in Cancer Immunotherapy: Therapies, Trials, and Challenges.Cancers (Basel). 2025 Feb 27;17(5):821. doi: 10.3390/cancers17050821. Cancers (Basel). 2025. PMID: 40075668 Free PMC article. Review.
-
OncomiRs as noncoding RNAs having functions in cancer: Their role in immune suppression and clinical implications.Front Immunol. 2022 Sep 16;13:913951. doi: 10.3389/fimmu.2022.913951. eCollection 2022. Front Immunol. 2022. PMID: 36189271 Free PMC article. Review.
References
-
- Shah V, Shah J. Recent trends in targeting miRNAs for cancer therapy. J Pharm Pharmacol. 2020. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

