Customized optical mapping by CRISPR-Cas9 mediated DNA labeling with multiple sgRNAs
- PMID: 33231685
- PMCID: PMC7826249
- DOI: 10.1093/nar/gkaa1088
Customized optical mapping by CRISPR-Cas9 mediated DNA labeling with multiple sgRNAs
Abstract
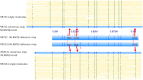
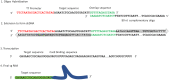
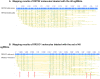
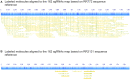
Whole-genome mapping technologies have been developed as a complementary tool to provide scaffolds for genome assembly and structural variation analysis (1,2). We recently introduced a novel DNA labeling strategy based on a CRISPR-Cas9 genome editing system, which can target any 20bp sequences. The labeling strategy is specifically useful in targeting repetitive sequences, and sequences not accessible to other labeling methods. In this report, we present customized mapping strategies that extend the applications of CRISPR-Cas9 DNA labeling. We first design a CRISPR-Cas9 labeling strategy to interrogate and differentiate the single allele differences in NGG protospacer adjacent motifs (PAM sequence). Combined with sequence motif labeling, we can pinpoint the single-base differences in highly conserved sequences. In the second strategy, we design mapping patterns across a genome by selecting sets of specific single-guide RNAs (sgRNAs) for labeling multiple loci of a genomic region or a whole genome. By developing and optimizing a single tube synthesis of multiple sgRNAs, we demonstrate the utility of CRISPR-Cas9 mapping with 162 sgRNAs targeting the 2Mb Haemophilus influenzae chromosome. These CRISPR-Cas9 mapping approaches could be particularly useful for applications in defining long-distance haplotypes and pinpointing the breakpoints in large structural variants in complex genomes and microbial mixtures.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

References
-
- Samad A., Huff E.J., Cai W.W., Schwartz D.C.. Optical mapping - a novel, single-molecule approach to genomic analysis. Genome Res. 1995; 5:1–4. - PubMed
-
- Gillett W., Hanks L., Wong G.K.-S., Yu J., Lim R., Olson M.V.J.G.. Assembly of high-resolution restriction maps based on multiple complete digests of a redundant set of overlapping clones. Genomics. 1996; 33:389–408. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

