Deciphering Streptococcal Biofilms
- PMID: 33233415
- PMCID: PMC7700319
- DOI: 10.3390/microorganisms8111835
Deciphering Streptococcal Biofilms
Abstract
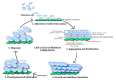
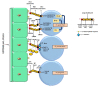
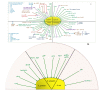
Streptococci are a diverse group of bacteria, which are mostly commensals but also cause a considerable proportion of life-threatening infections. They colonize many different host niches such as the oral cavity, the respiratory, gastrointestinal, and urogenital tract. While these host compartments impose different environmental conditions, many streptococci form biofilms on mucosal membranes facilitating their prolonged survival. In response to environmental conditions or stimuli, bacteria experience profound physiologic and metabolic changes during biofilm formation. While investigating bacterial cells under planktonic and biofilm conditions, various genes have been identified that are important for the initial step of biofilm formation. Expression patterns of these genes during the transition from planktonic to biofilm growth suggest a highly regulated and complex process. Biofilms as a bacterial survival strategy allow evasion of host immunity and protection against antibiotic therapy. However, the exact mechanisms by which biofilm-associated bacteria cause disease are poorly understood. Therefore, advanced molecular techniques are employed to identify gene(s) or protein(s) as targets for the development of antibiofilm therapeutic approaches. We review our current understanding of biofilm formation in different streptococci and how biofilm production may alter virulence-associated characteristics of these species. In addition, we have summarized the role of surface proteins especially pili proteins in biofilm formation. This review will provide an overview of strategies which may be exploited for developing novel approaches against biofilm-related streptococcal infections.
Keywords: antibiotic therapy; biofilm; opportunistic pathogen; planktonic; quorum sensing (QS); streptococci.
Conflict of interest statement
The authors declare no conflict of interest
Figures
Similar articles
-
Broad-spectrum quorum sensing and biofilm inhibition by green tea against gram-negative pathogenic bacteria: Deciphering the role of phytocompounds through molecular modelling.Microb Pathog. 2019 Jan;126:379-392. doi: 10.1016/j.micpath.2018.11.030. Epub 2018 Nov 23. Microb Pathog. 2019. PMID: 30476580
-
Piper betle and its bioactive metabolite phytol mitigates quorum sensing mediated virulence factors and biofilm of nosocomial pathogen Serratia marcescens in vitro.J Ethnopharmacol. 2016 Dec 4;193:592-603. doi: 10.1016/j.jep.2016.10.017. Epub 2016 Oct 6. J Ethnopharmacol. 2016. PMID: 27721053
-
Quorum sensing in streptococcal biofilm formation.Trends Microbiol. 2005 Jan;13(1):3-6. doi: 10.1016/j.tim.2004.11.009. Trends Microbiol. 2005. PMID: 15639624 Review.
-
Emergent Properties in Streptococcus mutans Biofilms Are Controlled through Adhesion Force Sensing by Initial Colonizers.mBio. 2019 Sep 10;10(5):e01908-19. doi: 10.1128/mBio.01908-19. mBio. 2019. PMID: 31506311 Free PMC article.
-
Quorum sensing in biofilms--how to destroy the bacterial citadels or their cohesion/power?Anaerobe. 2011 Dec;17(6):280-5. doi: 10.1016/j.anaerobe.2011.03.023. Epub 2011 Apr 8. Anaerobe. 2011. PMID: 21497662 Review.
Cited by
-
The Role of Rifampin in Prosthetic Joint Infections: Efficacy, Challenges, and Clinical Evidence.Antibiotics (Basel). 2024 Dec 17;13(12):1223. doi: 10.3390/antibiotics13121223. Antibiotics (Basel). 2024. PMID: 39766613 Free PMC article. Review.
-
Exploring Condition-Specific Variability in the Ureteral Stent Microbiome.Pathogens. 2024 Oct 29;13(11):942. doi: 10.3390/pathogens13110942. Pathogens. 2024. PMID: 39599495 Free PMC article.
-
Amyloid Aggregates Are Localized to the Nonadherent Detached Fraction of Aging Streptococcus mutans Biofilms.Microbiol Spectr. 2022 Aug 31;10(4):e0166122. doi: 10.1128/spectrum.01661-22. Epub 2022 Aug 11. Microbiol Spectr. 2022. PMID: 35950854 Free PMC article.
-
Antibiotic resistance, biofilm formation, and virulence genes of Streptococcus agalactiae serotypes of Indian origin.BMC Microbiol. 2023 Jul 5;23(1):176. doi: 10.1186/s12866-023-02877-y. BMC Microbiol. 2023. PMID: 37407919 Free PMC article.
-
The First Case of Streptococcus sinensis Endocarditis in Italy: Case Presentation and Systematic Literature Review.Medicina (Kaunas). 2024 Dec 2;60(12):1991. doi: 10.3390/medicina60121991. Medicina (Kaunas). 2024. PMID: 39768871 Free PMC article.
References
-
- Garrett T., Bhakoo M., Zhang Z. Bacterial adhesion and biofilms on surfaces. Prog. Nat. Sci. 2008;18 doi: 10.1016/j.pnsc.2008.04.001. - DOI
Publication types
LinkOut - more resources
Full Text Sources

