Method comparison of targeted influenza A virus typing and whole-genome sequencing from respiratory specimens of companion animals
- PMID: 33234046
- PMCID: PMC7953085
- DOI: 10.1177/1040638720933875
Method comparison of targeted influenza A virus typing and whole-genome sequencing from respiratory specimens of companion animals
Abstract
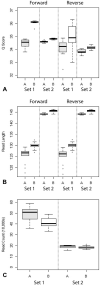
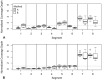
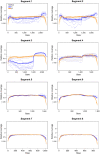
Epidemics of H3N8 and H3N2 influenza A viruses (IAVs) in dogs, along with recognition of spillover infections from IAV strains typically found in humans or other animals, have emphasized the importance of efficient laboratory testing. Given the lack of active IAV surveillance or immunization requirements for dogs, cats, or horses imported into the United States, serotype prediction and whole-genome sequencing of positive specimens detected at veterinary diagnostic laboratories are also needed. The conserved sequences at the ends of the viral genome segments facilitate universal amplification of all segments of viral genomes directly from respiratory specimens. Although several methods for genomic analysis have been reported, no optimization focusing on companion animal strains has been described, to our knowledge. We compared 2 sets of published universal amplification primers using 26 IAV-positive specimens from dogs, horses, and a cat. Libraries prepared from the resulting amplicons were sequenced using Illumina chemistry, and reference-based assemblies were generated from the data produced by both methods. Although both methods produced high-quality data, coverage profiles and base calling differed between the 2 methods. The sequence data were also used to identify the subtype of the IAV strains sequenced and then compared to standard PCR assays for neuraminidase types N2 and N8.
Keywords: M-RT-PCR; canine; equine; feline; influenza A virus; influenza matrix; multi-segment PCR; neuraminidase; reverse-transcription PCR; single-nucleotide polymorphism; typing; whole-genome sequencing.
Conflict of interest statement
Figures


Similar articles
-
An efficient genome sequencing method for equine influenza [H3N8] virus reveals a new polymorphism in the PA-X protein.Virol J. 2014 Sep 2;11:159. doi: 10.1186/1743-422X-11-159. Virol J. 2014. PMID: 25183201 Free PMC article.
-
Canine and Feline Influenza.Cold Spring Harb Perspect Med. 2021 Jan 4;11(1):a038562. doi: 10.1101/cshperspect.a038562. Cold Spring Harb Perspect Med. 2021. PMID: 31871238 Free PMC article. Review.
-
Design and testing of multiplex RT-PCR primers for the rapid detection of influenza A virus genomic segments: Application to equine influenza virus.J Virol Methods. 2016 Feb;228:114-22. doi: 10.1016/j.jviromet.2015.11.012. Epub 2015 Dec 4. J Virol Methods. 2016. PMID: 26655588
-
Phylogenetic Analysis and Characterization of a Sporadic Isolate of Equine Influenza A H3N8 from an Unvaccinated Horse in 2015.Viruses. 2018 Jan 11;10(1):31. doi: 10.3390/v10010031. Viruses. 2018. PMID: 29324680 Free PMC article.
-
H3N8 and H3N2 Canine Influenza Viruses: Understanding These New Viruses in Dogs.Vet Clin North Am Small Anim Pract. 2019 Jul;49(4):643-649. doi: 10.1016/j.cvsm.2019.02.005. Epub 2019 Apr 4. Vet Clin North Am Small Anim Pract. 2019. PMID: 30956002 Review.
Cited by
-
Whole-Genome Analysis of Influenza A(H3N2) and B/Victoria Viruses Detected in Myanmar during the COVID-19 Pandemic in 2021.Viruses. 2023 Feb 20;15(2):583. doi: 10.3390/v15020583. Viruses. 2023. PMID: 36851797 Free PMC article.
-
IAVCP (Influenza A Virus Consensus and Phylogeny): Automatic Identification of the Genomic Sequence of the Influenza A Virus from High-Throughput Sequencing Data.Viruses. 2024 May 29;16(6):873. doi: 10.3390/v16060873. Viruses. 2024. PMID: 38932165 Free PMC article.
-
Understanding the divergent evolution and epidemiology of H3N8 influenza viruses in dogs and horses.Virus Evol. 2023 Aug 18;9(2):vead052. doi: 10.1093/ve/vead052. eCollection 2023. Virus Evol. 2023. PMID: 37692894 Free PMC article.
-
Descriptive epidemiology and phylogenetic analysis of highly pathogenic avian influenza H5N1 clade 2.3.4.4b in British Columbia (B.C.) and the Yukon, Canada, September 2022 to June 2023.Emerg Microbes Infect. 2024 Dec;13(1):2392667. doi: 10.1080/22221751.2024.2392667. Epub 2024 Sep 22. Emerg Microbes Infect. 2024. PMID: 39143912 Free PMC article.
-
High pathogenicity avian influenza (HPAI) H7N6 virus detected in New Zealand poultry.Microbiol Resour Announc. 2025 Jun 12;14(6):e0008825. doi: 10.1128/mra.00088-25. Epub 2025 Apr 29. Microbiol Resour Announc. 2025. PMID: 40298424 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

