The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets
- PMID: 33237311
- PMCID: PMC7779004
- DOI: 10.1093/nar/gkaa1074
The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets
Erratum in
-
Correction to 'The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets'.Nucleic Acids Res. 2021 Oct 11;49(18):10800. doi: 10.1093/nar/gkab835. Nucleic Acids Res. 2021. PMID: 34530444 Free PMC article. No abstract available.
Abstract
Cellular life depends on a complex web of functional associations between biomolecules. Among these associations, protein-protein interactions are particularly important due to their versatility, specificity and adaptability. The STRING database aims to integrate all known and predicted associations between proteins, including both physical interactions as well as functional associations. To achieve this, STRING collects and scores evidence from a number of sources: (i) automated text mining of the scientific literature, (ii) databases of interaction experiments and annotated complexes/pathways, (iii) computational interaction predictions from co-expression and from conserved genomic context and (iv) systematic transfers of interaction evidence from one organism to another. STRING aims for wide coverage; the upcoming version 11.5 of the resource will contain more than 14 000 organisms. In this update paper, we describe changes to the text-mining system, a new scoring-mode for physical interactions, as well as extensive user interface features for customizing, extending and sharing protein networks. In addition, we describe how to query STRING with genome-wide, experimental data, including the automated detection of enriched functionalities and potential biases in the user's query data. The STRING resource is available online, at https://string-db.org/.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
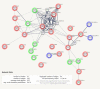
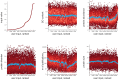
References
-
- Barabasi A.L., Oltvai Z.N.. Network biology: understanding the cell's functional organization. Nat. Rev. Genet. 2004; 5:101–113. - PubMed
-
- Hu J.X., Thomas C.E., Brunak S.. Network biology concepts in complex disease comorbidities. Nat. Rev. Genet. 2016; 17:615–629. - PubMed
-
- Conte F., Fiscon G., Licursi V., Bizzarri D., D’Anto T., Farina L., Paci P.. A paradigm shift in medicine: A comprehensive review of network-based approaches. Biochim. Biophys. Acta Gene Regul. Mech. 2020; 1863:194416. - PubMed
-
- Cowen L., Ideker T., Raphael B.J., Sharan R.. Network propagation: a universal amplifier of genetic associations. Nat. Rev. Genet. 2017; 18:551–562. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

