Hetero-oligomerization of Rho and Ras GTPases Connects GPCR Activation to mTORC2-AKT Signaling
- PMID: 33238110
- PMCID: PMC7716479
- DOI: 10.1016/j.celrep.2020.108427
Hetero-oligomerization of Rho and Ras GTPases Connects GPCR Activation to mTORC2-AKT Signaling
Abstract
The activation of G-protein-coupled receptors (GPCRs) leads to the activation of mTORC2 in cell migration and metabolism. However, the mechanism that links GPCRs to mTORC2 remains unknown. Here, using Dictyostelium cells, we show that GPCR-mediated chemotactic stimulation induces hetero-oligomerization of phosphorylated GDP-bound Rho GTPase and GTP-bound Ras GTPase in directed cell migration. The Rho-Ras hetero-oligomers directly and specifically stimulate mTORC2 activity toward AKT in cells and after biochemical reconstitution using purified proteins in vitro. The Rho-Ras hetero-oligomers do not activate ERK/MAPK, another kinase that functions downstream of GPCRs and Ras. Human KRas4B functionally replace Dictyostelium Ras in mTORC2 activation. In contrast to GDP-Rho, GTP-Rho antagonizes mTORC2-AKT signaling by inhibiting the oligomerization of GDP-Rho with GTP-Ras. These data reveal that GPCR-stimulated hetero-oligomerization of Rho and Ras provides a critical regulatory step that controls mTORC2-AKT signaling.
Keywords: AKT; Dictyostelium; G protein-coupled receptors; KRas; Rho; cell migration; mTORC2; small GTPases.
Copyright © 2020 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration Of Interests The authors declare no competing interests.
Figures





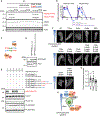

Similar articles
-
Phosphorylated Rho-GDP directly activates mTORC2 kinase towards AKT through dimerization with Ras-GTP to regulate cell migration.Nat Cell Biol. 2019 Jul;21(7):867-878. doi: 10.1038/s41556-019-0348-8. Epub 2019 Jul 1. Nat Cell Biol. 2019. PMID: 31263268 Free PMC article.
-
KARATE: PKA-induced KRAS4B-RHOA-mTORC2 supercomplex phosphorylates AKT in insulin signaling and glucose homeostasis.Mol Cell. 2021 Nov 18;81(22):4622-4634.e8. doi: 10.1016/j.molcel.2021.09.001. Epub 2021 Sep 21. Mol Cell. 2021. PMID: 34551282
-
Cyclic AMP-dependent and Epac-mediated activation of R-Ras by G protein-coupled receptors leads to phospholipase D stimulation.J Biol Chem. 2006 Aug 4;281(31):21837-21847. doi: 10.1074/jbc.M604156200. Epub 2006 Jun 5. J Biol Chem. 2006. PMID: 16754664
-
Regulation of phosphorylation pathways by p21 GTPases. The p21 Ras-related Rho subfamily and its role in phosphorylation signalling pathways.Eur J Biochem. 1996 Dec 1;242(2):171-85. doi: 10.1111/j.1432-1033.1996.0171r.x. Eur J Biochem. 1996. PMID: 8973630 Review.
-
G protein-coupled receptor oligomerization: implications for G protein activation and cell signaling.Circ Res. 2004 Jan 9;94(1):17-27. doi: 10.1161/01.RES.0000110420.68526.19. Circ Res. 2004. PMID: 14715532 Review.
Cited by
-
Inhibition of FNDC1 suppresses gastric cancer progression by interfering with Gβγ-VEGFR2 complex formation.iScience. 2023 Aug 12;26(9):107534. doi: 10.1016/j.isci.2023.107534. eCollection 2023 Sep 15. iScience. 2023. PMID: 37670789 Free PMC article.
-
FilGAP regulates tumor growth in Glioma through the regulation of mTORC1 and mTORC2.Sci Rep. 2023 Dec 8;13(1):20956. doi: 10.1038/s41598-023-47892-1. Sci Rep. 2023. PMID: 38065968 Free PMC article.
-
Regulation of the Actin Cytoskeleton via Rho GTPase Signalling in Dictyostelium and Mammalian Cells: A Parallel Slalom.Cells. 2021 Jun 24;10(7):1592. doi: 10.3390/cells10071592. Cells. 2021. PMID: 34202767 Free PMC article. Review.
-
The Crossroads between RAS and RHO Signaling Pathways in Cellular Transformation, Motility and Contraction.Genes (Basel). 2021 May 27;12(6):819. doi: 10.3390/genes12060819. Genes (Basel). 2021. PMID: 34071831 Free PMC article. Review.
-
Flexibility and Adaptation of Cancer Cells in a Heterogenous Metabolic Microenvironment.Int J Mol Sci. 2021 Feb 2;22(3):1476. doi: 10.3390/ijms22031476. Int J Mol Sci. 2021. PMID: 33540663 Free PMC article. Review.
References
-
- Ambrogio C, Kohler J, Zhou ZW, Wang H, Paranal R, Li J, Capelletti M, Caffarra C, Li S, Lv Q, et al. (2018). KRAS Dimerization Impacts MEK Inhibitor Sensitivity and Oncogenic Activity of Mutant KRAS. Cell 172, 857–868. - PubMed
-
- Caunt CJ, Sale MJ, Smith PD, and Cook SJ (2015). MEK1 and MEK2 inhibitors and cancer therapy: the long and winding road. Nat. Rev. Cancer 15, 577–592. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

