Biased localization of actin binding proteins by actin filament conformation
- PMID: 33239610
- PMCID: PMC7688639
- DOI: 10.1038/s41467-020-19768-9
Biased localization of actin binding proteins by actin filament conformation
Abstract
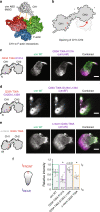
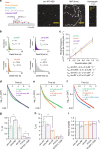
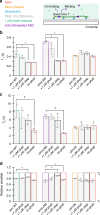
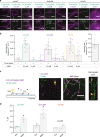
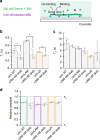
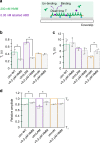
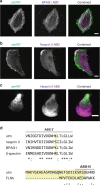
The assembly of actin filaments into distinct cytoskeletal structures plays a critical role in cell physiology, but how proteins localize differentially to these structures within a shared cytoplasm remains unclear. Here, we show that the actin-binding domains of accessory proteins can be sensitive to filament conformational changes. Using a combination of live cell imaging and in vitro single molecule binding measurements, we show that tandem calponin homology domains (CH1-CH2) can be mutated to preferentially bind actin networks at the front or rear of motile cells. We demonstrate that the binding kinetics of CH1-CH2 domain mutants varies as actin filament conformation is altered by perturbations that include stabilizing drugs and other binding proteins. These findings suggest that conformational changes of actin filaments in cells could help to direct accessory binding proteins to different actin cytoskeletal structures through a biophysical feedback loop.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Steric regulation of tandem calponin homology domain actin-binding affinity.Mol Biol Cell. 2019 Dec 15;30(26):3112-3122. doi: 10.1091/mbc.E19-06-0317. Epub 2019 Nov 6. Mol Biol Cell. 2019. PMID: 31693446 Free PMC article.
-
The N- and C-Terminal Domains Differentially Contribute to the Structure and Function of Dystrophin and Utrophin Tandem Calponin-Homology Domains.Biochemistry. 2015 Nov 24;54(46):6942-50. doi: 10.1021/acs.biochem.5b00969. Epub 2015 Nov 13. Biochemistry. 2015. PMID: 26516677
-
The actin binding affinity of the utrophin tandem calponin-homology domain is primarily determined by its N-terminal domain.Biochemistry. 2014 Mar 25;53(11):1801-9. doi: 10.1021/bi500149q. Epub 2014 Mar 14. Biochemistry. 2014. PMID: 24628267
-
Biophysics of actin filament severing by cofilin.FEBS Lett. 2013 Apr 17;587(8):1215-9. doi: 10.1016/j.febslet.2013.01.062. Epub 2013 Feb 5. FEBS Lett. 2013. PMID: 23395798 Free PMC article. Review.
-
The state of the filament.EMBO Rep. 2005 Mar;6(3):220-6. doi: 10.1038/sj.embor.7400363. EMBO Rep. 2005. PMID: 15741975 Free PMC article. Review.
Cited by
-
The open to closed D-loop conformational switch determines length in filopodia-like actin bundles.Biochem J. 2024 Dec 23;481(24):1977-1995. doi: 10.1042/BCJ20240367. Biochem J. 2024. PMID: 39621444 Free PMC article.
-
Myosin 2 - A general contractor for the cytoskeleton.Curr Opin Cell Biol. 2025 Jun;94:102522. doi: 10.1016/j.ceb.2025.102522. Epub 2025 May 3. Curr Opin Cell Biol. 2025. PMID: 40319507 Review.
-
Lymphocyte cytosolic protein 1 (L-plastin) I232F mutation impairs granulocytic proliferation and causes neutropenia.Blood Adv. 2022 Apr 26;6(8):2581-2594. doi: 10.1182/bloodadvances.2021006398. Blood Adv. 2022. PMID: 34991157 Free PMC article.
-
Multiple Mechanisms to Regulate Actin Functions: "Fundamental" Versus Lineage-Specific Mechanisms and Hierarchical Relationships.Biomolecules. 2025 Feb 13;15(2):279. doi: 10.3390/biom15020279. Biomolecules. 2025. PMID: 40001582 Free PMC article. Review.
-
Structural insights into actin isoforms.Elife. 2023 Feb 15;12:e82015. doi: 10.7554/eLife.82015. Elife. 2023. PMID: 36790143 Free PMC article.
References
-
- Harris AR, Jreij P, Fletcher DA. Mechanotransduction by the actin cytoskeleton: converting mechanical stimuli into biochemical signals. Annu. Rev. Biophys. 2018;47:617–631. doi: 10.1146/annurev-biophys-070816-033547. - DOI

