Mesenchymal Transformation: The Rosetta Stone of Glioblastoma Pathogenesis and Therapy Resistance
- PMID: 33240762
- PMCID: PMC7675056
- DOI: 10.1002/advs.202002015
Mesenchymal Transformation: The Rosetta Stone of Glioblastoma Pathogenesis and Therapy Resistance
Abstract
Despite decades of research, glioblastoma (GBM) remains invariably fatal among all forms of cancers. The high level of inter- and intratumoral heterogeneity along with its biological location, the brain, are major barriers against effective treatment. Molecular and single cell analysis identifies different molecular subtypes with varying prognosis, while multiple subtypes can reside in the same tumor. Cellular plasticity among different subtypes in response to therapies or during recurrence adds another hurdle in the treatment of GBM. This phenotypic shift is induced and sustained by activation of several pathways within the tumor itself, or microenvironmental factors. In this review, the dynamic nature of cellular shifts in GBM and how the tumor (immune) microenvironment shapes this process leading to therapeutic resistance, while highlighting emerging tools and approaches to study this dynamic double-edged sword are discussed.
Keywords: clinical outcome; glioblastoma (GBM); mesenchymal transition; molecular subtypes; therapy responses; tumor microenvironment.
© 2020 The Authors. Published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

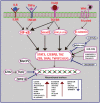


References
-
- Filbin M., Sturm D., Semin. Neurol. 2018, 38, 121. - PubMed
-
- Buckner J. C., Brown P. D., O'Neill B. P., Meyer F. B., Wetmore C. J., Uhm J. H., Mayo Clin. Proc. 2007, 82, 1271. - PubMed
-
- Stupp R., Hegi M. E., Mason W. P., Van Den Bent M. J., Taphoorn M. J., Janzer R. C., Ludwin S. K., Allgeier A., Fisher B., Belanger K., Hau P., Brandes A. A., Gijtenbeek J., Marosi C., Vecht C. J., Mokhtari K., Wesseling P., Villa S., Eisenhauer E., Gorlia T., Weller M., Lacombe D., Cairncross J. G., Mirimanoff R.‐O., Lancet Oncol. 2009, 10, 459. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
