DNA Polymerases at the Eukaryotic Replication Fork Thirty Years after: Connection to Cancer
- PMID: 33255191
- PMCID: PMC7760166
- DOI: 10.3390/cancers12123489
DNA Polymerases at the Eukaryotic Replication Fork Thirty Years after: Connection to Cancer
Erratum in
-
Correction: Pavlov, Y.I., et al. DNA Polymerases at the Eukaryotic Replication Fork Thirty Years After: Connection to Cancer. Cancers 2020, 12, 3489.Cancers (Basel). 2021 Feb 26;13(5):969. doi: 10.3390/cancers13050969. Cancers (Basel). 2021. PMID: 33653017 Free PMC article.
Abstract
Recent studies on tumor genomes revealed that mutations in genes of replicative DNA polymerases cause a predisposition for cancer by increasing genome instability. The past 10 years have uncovered exciting details about the structure and function of replicative DNA polymerases and the replication fork organization. The principal idea of participation of different polymerases in specific transactions at the fork proposed by Morrison and coauthors 30 years ago and later named "division of labor," remains standing, with an amendment of the broader role of polymerase δ in the replication of both the lagging and leading DNA strands. However, cancer-associated mutations predominantly affect the catalytic subunit of polymerase ε that participates in leading strand DNA synthesis. We analyze how new findings in the DNA replication field help elucidate the polymerase variants' effects on cancer.
Keywords: DNA polymerases; cancer predisposition; mutation rates; proofreading exonucleases; replication fidelity.
Conflict of interest statement
The authors declare no conflict of interest. There is no role of funding agencies in the choice of review, writing of the manuscript, or publishing results.
Figures

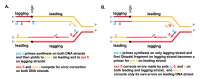

 ) is present in each of the four pol complexes.
) is present in each of the four pol complexes.
References
-
- Loeb L.A., Springgate C.F., Battula N. Errors in DNA replication as a basis of malignant changes. Cancer Res. 1974;34:2311–2321. - PubMed
-
- Tahirov T.H., Makarova K.S., Rogozin I.B., Pavlov Y.I., Koonin E.V. Evolution of DNA polymerases: An inactivated polymerase-exonuclease module in Pol epsilon and a chimeric origin of eukaryotic polymerases from two classes of archaeal ancestors. Biol. Direct. 2009;4:11. doi: 10.1186/1745-6150-4-11. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

