Electron Attachment Studies with the Potential Radiosensitizer 2-Nitrofuran
- PMID: 33255344
- PMCID: PMC7727711
- DOI: 10.3390/ijms21238906
Electron Attachment Studies with the Potential Radiosensitizer 2-Nitrofuran
Abstract
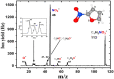
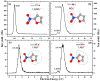
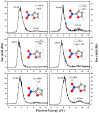
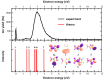
Nitrofurans belong to the class of drugs typically used as antibiotics or antimicrobials. The defining structural component is a furan ring with a nitro group attached. In the present investigation, electron attachment to 2-nitrofuran (C4H3NO3), which is considered as a potential radiosensitizer candidate for application in radiotherapy, has been studied in a crossed electron-molecular beams experiment. The present results indicate that low-energy electrons with kinetic energies of about 0-12 eV effectively decompose the molecule. In total, twelve fragment anions were detected within the detection limit of the apparatus, as well as the parent anion of 2-nitrofuran. One major resonance region of ≈0-5 eV is observed in which the most abundant anions NO2-, C4H3O-, and C4H3NO3- are detected. The experimental results are supported by ab initio calculations of electronic states in the resulting anion, thermochemical thresholds, connectivity between electronic states of the anion, and reactivity analysis in the hot ground state.
Keywords: electron attachment; fragmentation; low-energy electron; nitrofuran; radiosensitizer; reduction.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Low-Energy Electron Induced Reactions in Metronidazole at Different Solvation Conditions.Pharmaceuticals (Basel). 2022 Jun 2;15(6):701. doi: 10.3390/ph15060701. Pharmaceuticals (Basel). 2022. PMID: 35745620 Free PMC article.
-
Dissociative Electron Attachment to 5-Iodo-4-thio-2'-deoxyuridine: A Potential Radiosensitizer of Hypoxic Cells.J Phys Chem Lett. 2023 Oct 12;14(40):8948-8955. doi: 10.1021/acs.jpclett.3c02219. Epub 2023 Sep 28. J Phys Chem Lett. 2023. PMID: 37769041 Free PMC article.
-
Formation of negative and positive ions in the radiosensitizer nimorazole upon low-energy electron collisions.J Chem Phys. 2021 Feb 21;154(7):074306. doi: 10.1063/5.0040045. J Chem Phys. 2021. PMID: 33607883
-
Ring-Selective Fragmentation in the Tirapazamine Molecule upon Low-Energy Electron Attachment.Int J Mol Sci. 2021 Mar 19;22(6):3159. doi: 10.3390/ijms22063159. Int J Mol Sci. 2021. PMID: 33808887 Free PMC article.
-
Clustered DNA Damages induced by 0.5 to 30 eV Electrons.Int J Mol Sci. 2019 Jul 31;20(15):3749. doi: 10.3390/ijms20153749. Int J Mol Sci. 2019. PMID: 31370253 Free PMC article. Review.
Cited by
-
The Physical Chemistry and Chemical Physics (PCCP) Section of the International Journal of Molecular Sciences in Its Publications: The First 300 Thematic Articles in the First 3 Years.Int J Mol Sci. 2021 Dec 27;23(1):241. doi: 10.3390/ijms23010241. Int J Mol Sci. 2021. PMID: 35008667 Free PMC article.
-
Electron-Induced Decomposition of Uracil-5-yl O-(N,N-dimethylsulfamate): Role of Methylation in Molecular Stability.Int J Mol Sci. 2021 Feb 26;22(5):2344. doi: 10.3390/ijms22052344. Int J Mol Sci. 2021. PMID: 33652878 Free PMC article.
-
Low-Energy Electron Induced Reactions in Metronidazole at Different Solvation Conditions.Pharmaceuticals (Basel). 2022 Jun 2;15(6):701. doi: 10.3390/ph15060701. Pharmaceuticals (Basel). 2022. PMID: 35745620 Free PMC article.
-
Electron Attachment to 5-Fluorouracil: The Role of Hydrogen Fluoride in Dissociation Chemistry.Int J Mol Sci. 2022 Jul 28;23(15):8325. doi: 10.3390/ijms23158325. Int J Mol Sci. 2022. PMID: 35955461 Free PMC article.
-
Decomposition of triazole and 3-nitrotriazole upon low-energy electron attachment.Phys Chem Chem Phys. 2023 May 24;25(20):13892-13901. doi: 10.1039/d3cp01162c. Phys Chem Chem Phys. 2023. PMID: 37183636 Free PMC article.
References
-
- Gorfinkiel J.D., Ptasinska S. Electron scattering from molecules and molecular aggregates of biological relevance. J. Phys. B At. Mol. Opt. Phys. 2017;50 doi: 10.1088/1361-6455/aa8572. - DOI
-
- Baccarelli I., Bald I., Gianturco F.A., Illenberger E., Kopyra J. Electron-induced damage of DNA and its components: Experiments and theoretical models. Phys. Rep. 2011;508:1–44. doi: 10.1016/j.physrep.2011.06.004. - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

