Integrin-Linked Kinase Is a Novel Therapeutic Target in Ovarian Cancer
- PMID: 33256002
- PMCID: PMC7712057
- DOI: 10.3390/jpm10040246
Integrin-Linked Kinase Is a Novel Therapeutic Target in Ovarian Cancer
Abstract
Objective: The objective of this study is to identify and validate novel therapeutic target(s) in ovarian cancer.
Background: Development of targeted therapeutics in ovarian cancer has been limited by molecular heterogeneity. Although gene expression datasets are available, most of them lack appropriate pair-matched controls to define the alterations that result in the transformation of normal ovarian cells to cancerous cells.
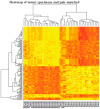
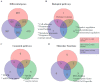
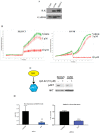
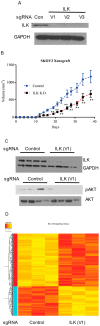
Methods: We used microarray to compare the gene expression of treatment-naïve ovarian cancer tissue samples to pair-matched normal adjacent ovarian tissue from 24 patients. Ingenuity Pathway Analysis (IPA) was used to identify target pathways for further analysis. Integrin-linked kinase (ILK) expression in SKOV3 and OV90 cells was determined using Western blot. ILK was knocked down using CRISPR/Cas9 constructs. Subcutaneous xenograft study to determine the effect of ILK knockdown on tumor growth was performed in NOD SCID gamma mice.
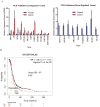
Results: Significant upregulation of the ILK pathway was identified in 22 of the 24 cancer specimens, identifying it as a potential player that could contribute to the transformation of normal ovarian cells to cancerous cells. Knockdown of ILK in SKOV3 cells resulted in decreased cell proliferation and tumor growth, and inhibition of downstream kinase, AKT (protein kinase B). These results were further validated using an ILK-1 chemical inhibitor, compound 22.
Conclusion: Our initial findings validate ILK as a potential therapeutic target for molecular inhibition in ovarian cancer, which warrants further investigation.
Keywords: gene expression; integrin-linked kinase (ILK); microarray; ovarian cancer; sgRNA; xenograft.
Conflict of interest statement
No authors have any relevant conflict of interest to disclose.
Figures

References
-
- Sallinen H., Janhonen S., Pölönen P., Niskanen H., Liu O.H., Kivelä A., Hartikainen J.M., Anttila M., Heinäniemi M., Yla-Herttuala S., et al. Comparative transcriptome analysis of matched primary and distant metastatic ovarian carcinoma. BMC Cancer. 2019;19:1–11. doi: 10.1186/s12885-019-6339-0. - DOI - PMC - PubMed
-
- Ye X. Confluence analysis of multiple omics on platinum resistance of ovarian cancer. Eur. J. Gynaecol. Oncol. 2015;36 - PubMed
LinkOut - more resources
Full Text Sources

