Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis
- PMID: 33256112
- PMCID: PMC7760495
- DOI: 10.3390/cancers12123528
Epigenetic Inactivation of the Tumor Suppressor IRX1 Occurs Frequently in Lung Adenocarcinoma and Its Silencing Is Associated with Impaired Prognosis
Abstract
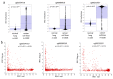
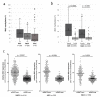
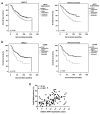
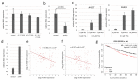
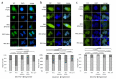
Iroquois homeobox (IRX) encodes members of homeodomain containing genes which are involved in development and differentiation. Since it has been reported that the IRX1 gene is localized in a lung cancer susceptibility locus, the epigenetic regulation and function of IRX1 was investigated in lung carcinogenesis. We observed frequent hypermethylation of the IRX1 promoter in non-small cell lung cancer (NSCLC) compared to small cell lung cancer (SCLC). Aberrant IRX1 methylation was significantly correlated with reduced IRX1 expression. In normal lung samples, the IRX1 promoter showed lower median DNA methylation levels (<10%) compared to primary adenocarcinoma (ADC, 22%) and squamous cell carcinoma (SQCC, 14%). A significant hypermethylation and downregulation of IRX1 was detected in ADC and SQCC compared to matching normal lung samples (p < 0.0001). Low IRX1 expression was significantly correlated with impaired prognosis of ADC patients (p = 0.001). Reduced survival probability was also associated with higher IRX1 promoter methylation (p = 0.02). Inhibition of DNA methyltransferase (DNMT) activity reactivated IRX1 expression in human lung cancer cell lines. Induced DNMT3A and EZH2 expression was correlated with downregulation of IRX1. On the cellular level, IRX1 exhibits nuclear localization and expression of IRX1 induced fragmented nuclei in cancer cells. Localization of IRX1 and induction of aberrant nuclei were dependent on the presence of the homeobox of IRX1. By data mining, we showed that IRX1 is negatively correlated with oncogenic pathways and IRX1 expression induces the proapoptotic regulator BAX. In conclusion, we report that IRX1 expression is significantly associated with improved survival probability of ADC patients. IRX1 hypermethylation may serve as molecular biomarker for ADC diagnosis and prognosis. Our data suggest that IRX1 acts as an epigenetically regulated tumor suppressor in the pathogenesis of lung cancer.
Keywords: DNA methylation; IRX; adenocarcinoma; apoptosis; epigenetic; homeobox; lung cancer; tumor suppressor.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript or in the decision to publish the results.
Figures


References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

