Susceptibility of Elizabethkingia spp. to commonly tested and novel antibiotics and concordance between broth microdilution and automated testing methods
- PMID: 33258923
- PMCID: PMC8600004
- DOI: 10.1093/jac/dkaa499
Susceptibility of Elizabethkingia spp. to commonly tested and novel antibiotics and concordance between broth microdilution and automated testing methods
Abstract
Objectives: We aimed to determine susceptibilities of Elizabethkingia spp. to 25 commonly tested and 8 novel antibiotics, and to compare the performance of different susceptibility testing methods.
Methods: Clinical isolates of Elizabethkingia spp., Chryseobacterium spp. and Flavobacterium spp. collected during 2002-18 (n = 210) in a nationwide surveillance programme in Taiwan were speciated by 16S rRNA sequencing. MICs were determined by broth microdilution. The broth microdilution results of 18 common antibiotics were compared with those obtained by the VITEK 2 automated system.
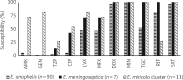
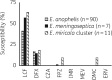
Results: Among the Elizabethkingia spp. identified (n = 108), Elizabethkingia anophelis was the most prevalent (n = 90), followed by Elizabethkingia meningoseptica (n = 7) and Elizabethkingia miricola cluster [E. miricola (n = 6), Elizabethkingia bruuniana (n = 3) and Elizabethkingia ursingii (n = 2)]. Most isolates were recovered from respiratory or blood specimens from hospitalized, elderly patients. PFGE showed two major and several minor E. anophelis clones. All isolates were resistant to nearly all the tested β-lactams. Doxycycline, minocycline and trimethoprim/sulfamethoxazole inhibited >90% of Elizabethkingia spp. Rifampin inhibited E. meningoseptica (100%) and E. anophelis (81.1%). Fluoroquinolones and tigecycline were active against E. meningoseptica and E. miricola cluster isolates. Novel antibiotics, including imipenem/relebactam, meropenem/vaborbactam, ceftazidime/avibactam, cefepime/zidebactam, delafloxacin, eravacycline and omadacycline were ineffective but lascufloxacin inhibited half of Elizabethkingia spp. The very major discrepancy rates of VITEK 2 were >1.5% for ciprofloxacin, moxifloxacin and vancomycin. Major discrepancy rates were >3% for amikacin, tigecycline, piperacillin/tazobactam and trimethoprim/sulfamethoxazole.
Conclusions: MDR, absence of standard interpretation criteria and poor intermethod concordance necessitate working guidelines to facilitate future research of emerging Elizabethkingia spp.
© The Author(s) 2020. Published by Oxford University Press on behalf of the British Society for Antimicrobial Chemotherapy. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures
References
-
- Nicholson AC, Gulvik CA, Whitney AM. et al. Revisiting the taxonomy of the genus Elizabethkingia using whole-genome sequencing, optical mapping, and MALDI-TOF, along with proposal of three novel Elizabethkingia species: Elizabethkingia bruuniana sp. nov., Elizabethkingia ursingii sp. nov., and Elizabethkingia occulta sp. nov. Antonie van Leeuwenhoek 2018; 111: 55–72. - PMC - PubMed
-
- Kim KK, Kim MK, Lim JH. et al. Transfer of Chryseobacterium meningosepticum and Chryseobacterium miricola to Elizabethkingia gen. nov. as Elizabethkingia meningoseptica comb. nov. and Elizabethkingia miricola comb. nov. Int J Syst Evol Microbiol 2005; 55: 1287–93. - PubMed
-
- Hsu MS, Liao CH, Huang YT. et al. Clinical features, antimicrobial susceptibilities, and outcomes of Elizabethkingia meningoseptica (Chryseobacterium meningosepticum) bacteremia at a medical center in Taiwan, 1999-2006. Eur J Clin Microbiol Infect Dis 2011; 30: 1271–8. - PubMed
-
- Li Y, Kawamura Y, Fujiwara N. et al. Chryseobacterium miricola sp. nov., a novel species isolated from condensation water of space station Mir. Syst Appl Microbiol 2003; 26: 523–8. - PubMed
-
- Kampfer P, Matthews H, Glaeser SP. et al. Elizabethkingia anophelis sp. nov., isolated from the midgut of the mosquito Anopheles gambiae. Int J Syst Evol Microbiol 2011; 61: 2670–5. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

