A Clustering Approach for Motif Discovery in ChIP-Seq Dataset
- PMID: 33267515
- PMCID: PMC7515331
- DOI: 10.3390/e21080802
A Clustering Approach for Motif Discovery in ChIP-Seq Dataset
Abstract
Chromatin immunoprecipitation combined with next-generation sequencing (ChIP-Seq) technology has enabled the identification of transcription factor binding sites (TFBSs) on a genome-wide scale. To effectively and efficiently discover TFBSs in the thousand or more DNA sequences generated by a ChIP-Seq data set, we propose a new algorithm named AP-ChIP. First, we set two thresholds based on probabilistic analysis to construct and further filter the cluster subsets. Then, we use Affinity Propagation (AP) clustering on the candidate cluster subsets to find the potential motifs. Experimental results on simulated data show that the AP-ChIP algorithm is able to make an almost accurate prediction of TFBSs in a reasonable time. Also, the validity of the AP-ChIP algorithm is tested on a real ChIP-Seq data set.
Keywords: ChIP-Seq; motif discovery; planted motif search; transcription factor binding sites.
Conflict of interest statement
The authors declare no conflict of interest.
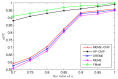
Figures
References
-
- Pevzner P.A., Sze S.H. ISMB. Volume 8. American Association for Artificial Intelligence; Menlo Park, CA, USA: 2000. Combinatorial approaches to finding subtle signals in DNA sequences; pp. 269–278. - PubMed
-
- Bailey T., Elkan C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994;2:28–36. - PubMed
-
- Chin F.Y., Leung H.C. Voting algorithms for discovering long motifs; Proceedings of the 3rd Asia-Pacific Bioinformatics Conference; Singapore. 17–21 January 2005; pp. 261–271.
Grants and funding
LinkOut - more resources
Full Text Sources