Molecular and evolutionary processes generating variation in gene expression
- PMID: 33268840
- PMCID: PMC7981258
- DOI: 10.1038/s41576-020-00304-w
Molecular and evolutionary processes generating variation in gene expression
Abstract
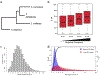
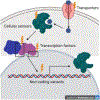
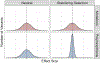
Heritable variation in gene expression is common within and between species. This variation arises from mutations that alter the form or function of molecular gene regulatory networks that are then filtered by natural selection. High-throughput methods for introducing mutations and characterizing their cis- and trans-regulatory effects on gene expression (particularly, transcription) are revealing how different molecular mechanisms generate regulatory variation, and studies comparing these mutational effects with variation seen in the wild are teasing apart the role of neutral and non-neutral evolutionary processes. This integration of molecular and evolutionary biology allows us to understand how the variation in gene expression we see today came to be and to predict how it is most likely to evolve in the future.
Figures
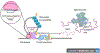
References
-
- Zheng W, Gianoulis TA, Karczewski KJ, Zhao H & Snyder M Regulatory variation within and between species. Annu. Rev. Genomics Hum. Genet 12, 327–346 (2011). - PubMed
-
- Kronforst MR et al. Unraveling the thread of nature’s tapestry: the genetics of diversity and convergence in animal pigmentation. Pigment Cell Melanoma Res. 25, 411–433 (2012). - PubMed
-
- Wessinger CA & Rausher MD Lessons from flower colour evolution on targets of selection. J. Exp. Bot 63, 5741–5749 (2012). - PubMed
-
- Deutschbauer AM & Davis RW Quantitative trait loci mapped to single-nucleotide resolution in yeast. Nat. Genet 37, 1333–1340 (2005). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

