TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
- PMID: 33273453
- PMCID: PMC7713251
- DOI: 10.1038/s41467-020-19877-5
TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
Abstract
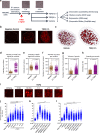
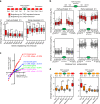
The Transforming Growth Factor-β (TGFβ) signaling pathway controls transcription by regulating enhancer activity. How TGFβ-regulated enhancers are selected and what chromatin changes are associated with TGFβ-dependent enhancers regulation are still unclear. Here we report that TGFβ treatment triggers fast and widespread increase in chromatin accessibility in about 80% of the enhancers of normal mouse mammary epithelial-gland cells, irrespective of whether they are activated, repressed or not regulated by TGFβ. This enhancer opening depends on both the canonical and non-canonical TGFβ pathways. Most TGFβ-regulated genes are located around enhancers regulated in the same way, often creating domains of several co-regulated genes that we term TGFβ regulatory domains (TRD). CRISPR-mediated inactivation of enhancers within TRDs impairs TGFβ-dependent regulation of all co-regulated genes, demonstrating that enhancer targeting is more promiscuous than previously anticipated. The area of TRD influence is restricted by topologically associating domains (TADs) borders, causing a bias towards co-regulation within TADs.
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Contribution of transposable elements and distal enhancers to evolution of human-specific features of interphase chromatin architecture in embryonic stem cells.Chromosome Res. 2018 Mar;26(1-2):61-84. doi: 10.1007/s10577-018-9571-6. Epub 2018 Jan 15. Chromosome Res. 2018. PMID: 29335803
-
Enhancer Reprogramming within Pre-existing Topologically Associated Domains Promotes TGF-β-Induced EMT and Cancer Metastasis.Mol Ther. 2020 Sep 2;28(9):2083-2095. doi: 10.1016/j.ymthe.2020.05.026. Epub 2020 Jun 1. Mol Ther. 2020. PMID: 32526202 Free PMC article.
-
JMJD3 intrinsically disordered region links the 3D-genome structure to TGFβ-dependent transcription activation.Nat Commun. 2022 Jun 7;13(1):3263. doi: 10.1038/s41467-022-30614-y. Nat Commun. 2022. PMID: 35672304 Free PMC article.
-
Evaluating Enhancer Function and Transcription.Annu Rev Biochem. 2020 Jun 20;89:213-234. doi: 10.1146/annurev-biochem-011420-095916. Epub 2020 Mar 20. Annu Rev Biochem. 2020. PMID: 32197056 Review.
-
Spatial genome organization, TGFβ, and biomolecular condensates: Do they talk during development?Bioessays. 2022 Dec;44(12):e2200145. doi: 10.1002/bies.202200145. Epub 2022 Oct 17. Bioessays. 2022. PMID: 36253122 Review.
Cited by
-
PHF13 epigenetically activates TGFβ driven epithelial to mesenchymal transition.Cell Death Dis. 2022 May 21;13(5):487. doi: 10.1038/s41419-022-04940-4. Cell Death Dis. 2022. PMID: 35597793 Free PMC article.
-
Telomemore enables single-cell analysis of cell cycle and chromatin condensation.Nucleic Acids Res. 2025 Jan 24;53(3):gkaf031. doi: 10.1093/nar/gkaf031. Nucleic Acids Res. 2025. PMID: 39878215 Free PMC article.
-
TGF-β-Upregulated Lnc-Nr6a1 Acts as a Reservoir of miR-181 and Mediates Assembly of a Glycolytic Complex.Noncoding RNA. 2022 Sep 15;8(5):62. doi: 10.3390/ncrna8050062. Noncoding RNA. 2022. PMID: 36136852 Free PMC article.
-
Excessive MYC Orchestrates Macrophages induced Chromatin Remodeling to Sustain Micropapillary-Patterned Malignancy in Lung Adenocarcinoma.Adv Sci (Weinh). 2025 Mar;12(12):e2403851. doi: 10.1002/advs.202403851. Epub 2025 Feb 3. Adv Sci (Weinh). 2025. PMID: 39899538 Free PMC article.
-
SMAD2 linker phosphorylation impacts overall survival, proliferation, TGFβ1-dependent gene expression and pluripotency-related proteins in NSCLC.Br J Cancer. 2025 Jul;133(1):52-65. doi: 10.1038/s41416-025-02970-1. Epub 2025 May 3. Br J Cancer. 2025. PMID: 40319202 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

