Cell-Type-Specific Gene Regulatory Networks Underlying Murine Neonatal Heart Regeneration at Single-Cell Resolution
- PMID: 33296652
- PMCID: PMC7774872
- DOI: 10.1016/j.celrep.2020.108472
Cell-Type-Specific Gene Regulatory Networks Underlying Murine Neonatal Heart Regeneration at Single-Cell Resolution
Erratum in
-
Cell-Type-Specific Gene Regulatory Networks Underlying Murine Neonatal Heart Regeneration at Single-Cell Resolution.Cell Rep. 2021 May 25;35(8):109211. doi: 10.1016/j.celrep.2021.109211. Cell Rep. 2021. PMID: 34038730 Free PMC article. No abstract available.
Abstract
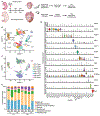
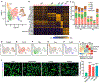
The adult mammalian heart has limited capacity for regeneration following injury, whereas the neonatal heart can readily regenerate within a short period after birth. Neonatal heart regeneration is orchestrated by multiple cell types intrinsic to the heart, as well as immune cells that infiltrate the heart after injury. To elucidate the transcriptional responses of the different cellular components of the mouse heart following injury, we perform single-cell RNA sequencing on neonatal hearts at various time points following myocardial infarction and couple the results with bulk tissue RNA-sequencing data collected at the same time points. Concomitant single-cell ATAC sequencing exposes underlying dynamics of open chromatin landscapes and regenerative gene regulatory networks of diverse cardiac cell types and reveals extracellular mediators of cardiomyocyte proliferation, angiogenesis, and fibroblast activation. Together, our data provide a transcriptional basis for neonatal heart regeneration at single-cell resolution and suggest strategies for enhancing cardiac function after injury.
Keywords: angiogenesis; cardiomyocyte proliferation; epicardial cells; fibrosis; injury response; myocardial infarction; open chromatin landscape; secreted factors; single cell ATAC-seq; single cell RNA-seq.
Copyright © 2020 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures





Comment in
-
Transcriptional atlas of mouse heart regeneration.Nat Rev Cardiol. 2021 Apr;18(4):231. doi: 10.1038/s41569-021-00509-4. Nat Rev Cardiol. 2021. PMID: 33441973 No abstract available.
References
-
- Abraityte A, Vinge LE, Askevold ET, Lekva T, Michelsen AE, Ranheim T, Alfsnes K, Fiane A, Aakhus S, Lunde IG, et al. (2017). Wnt5a is elevated in heart failure and affects cardiac fibroblast function. J. Mol. Med. (Berl.) 95, 767–777. - PubMed
-
- Alexander WS, Rakar S, Robb L, Farley A, Willson TA, Zhang JG, Hartley L, Kikuchi Y, Kojima T, Nomura H, et al. (1999). Suckling defect in mice lacking the soluble haemopoietin receptor NR6. Curr. Biol 9, 605–608. - PubMed
-
- Asp M, Giacomello S, Larsson L, Wu C, F€rth D, Qian X, Wardell E, Custodio J, Reimegård J, Salmén F, et al. (2019). A spatiotemporal organ-wide gene expression and cell atlas of the developing human heart. Cell 179, 1647–1660. e19. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

