Physiological Response of Corynebacterium glutamicum to Indole
- PMID: 33302489
- PMCID: PMC7764795
- DOI: 10.3390/microorganisms8121945
Physiological Response of Corynebacterium glutamicum to Indole
Abstract
The aromatic heterocyclic compound indole is widely spread in nature. Due to its floral odor indole finds application in dairy, flavor, and fragrance products. Indole is an inter- and intracellular signaling molecule influencing cell division, sporulation, or virulence in some bacteria that synthesize it from tryptophan by tryptophanase. Corynebacterium glutamicum that is used for the industrial production of amino acids including tryptophan lacks tryptophanase. To test if indole is metabolized by C. glutamicum or has a regulatory role, the physiological response to indole by this bacterium was studied. As shown by RNAseq analysis, indole, which inhibited growth at low concentrations, increased expression of genes involved in the metabolism of iron, copper, and aromatic compounds. In part, this may be due to iron reduction as indole was shown to reduce Fe3+ to Fe2+ in the culture medium. Mutants with improved tolerance to indole were selected by adaptive laboratory evolution. Among the mutations identified by genome sequencing, mutations in three transcriptional regulator genes were demonstrated to be causal for increased indole tolerance. These code for the regulator of iron homeostasis DtxR, the regulator of oxidative stress response RosR, and the hitherto uncharacterized Cg3388. Gel mobility shift analysis revealed that Cg3388 binds to the intergenic region between its own gene and the iolT2-rhcM2D2 operon encoding inositol uptake system IolT2, maleylacetate reductase, and catechol 1,2-dioxygenase. Increased RNA levels of rhcM2 in a cg3388 deletion strain indicated that Cg3388 acts as repressor. Indole, hydroquinone, and 1,2,4-trihydroxybenzene may function as inducers of the iolT2-rhcM2D2 operon in vivo as they interfered with DNA binding of Cg3388 at physiological concentrations in vitro. Cg3388 was named IhtR.
Keywords: Corynebacterium glutamicum; adaptive laboratory evolution; amino acids; aromatic compound catabolism; indole; iron homeostasis; oxidative stress.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data; in the writing of the manuscript or in the decision to publish the results.
Figures
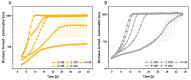





References
-
- Berger R.G., editor. Flavours and Fragrances: Chemistry, Bioprocessing and Sustainability. Springer; Berlin/Heidelberg, Germany: 2007. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

