Distinct polymorphisms in a single herpesvirus gene are capable of enhancing virulence and mediating vaccinal resistance
- PMID: 33306739
- PMCID: PMC7758048
- DOI: 10.1371/journal.ppat.1009104
Distinct polymorphisms in a single herpesvirus gene are capable of enhancing virulence and mediating vaccinal resistance
Abstract
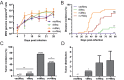
Modified-live herpesvirus vaccines are widely used in humans and animals, but field strains can emerge that have a higher virulence and break vaccinal protection. Since the introduction of the first vaccine in the 1970s, Marek's disease virus overcame the vaccine barrier by the acquisition of numerous genomic mutations. However, the evolutionary adaptations in the herpesvirus genome responsible for the vaccine breaks have remained elusive. Here, we demonstrate that point mutations in the multifunctional meq gene acquired during evolution can significantly alter virulence. Defined mutations found in highly virulent strains also allowed the virus to overcome innate cellular responses and vaccinal protection. Concomitantly, the adaptations in meq enhanced virus shedding into the environment, likely providing a selective advantage for the virus. Our study provides the first experimental evidence that few point mutations in a single herpesviral gene result in drastically increased virulence, enhanced shedding, and escape from vaccinal protection.
Conflict of interest statement
The authors have declared that no competing interests exist.
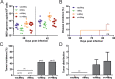
Figures



Similar articles
-
A Common Live-Attenuated Avian Herpesvirus Vaccine Expresses a Very Potent Oncogene.mSphere. 2019 Oct 9;4(5):e00658-19. doi: 10.1128/mSphere.00658-19. mSphere. 2019. PMID: 31597721 Free PMC article.
-
Recombinant Gallid herpesvirus 2 with interrupted meq genes confers safe and efficacious protection against virulent field strains.Vaccine. 2017 Aug 24;35(36):4695-4701. doi: 10.1016/j.vaccine.2017.07.048. Epub 2017 Jul 25. Vaccine. 2017. PMID: 28754487
-
Vaccinal control of Marek's disease: current challenges, and future strategies to maximize protection.Vet Immunol Immunopathol. 2006 Jul 15;112(1-2):78-86. doi: 10.1016/j.vetimm.2006.03.014. Epub 2006 May 8. Vet Immunol Immunopathol. 2006. PMID: 16682084 Review.
-
Differentiation between pathogenic serotype 1 isolates of Marek's disease virus and the Rispens CVI988 vaccine in Australia using real-time PCR and high resolution melt curve analysis.J Virol Methods. 2013 Jan;187(1):144-52. doi: 10.1016/j.jviromet.2012.09.018. Epub 2012 Oct 3. J Virol Methods. 2013. PMID: 23041147
-
History of the First-Generation Marek's Disease Vaccines: The Science and Little-Known Facts.Avian Dis. 2016 Dec;60(4):715-724. doi: 10.1637/11429-050216-Hist. Avian Dis. 2016. PMID: 27902902 Review.
Cited by
-
Telomeric repeats in the commercial SB-1 vaccine facilitate viral integration and contribute to vaccine efficacy.NPJ Vaccines. 2024 Aug 21;9(1):154. doi: 10.1038/s41541-024-00945-6. NPJ Vaccines. 2024. PMID: 39169010 Free PMC article.
-
In vivo imaging reveals novel replication sites of a highly oncogenic avian herpesvirus in chickens.PLoS Pathog. 2022 Aug 29;18(8):e1010745. doi: 10.1371/journal.ppat.1010745. eCollection 2022 Aug. PLoS Pathog. 2022. PMID: 36037230 Free PMC article.
-
Phylogenetic and Genomic Characterization of Whole Genome Sequences of Ocular Herpes Simplex Virus Type 1 Isolates Identifies Possible Virulence Determinants in Humans.Invest Ophthalmol Vis Sci. 2023 Jul 3;64(10):16. doi: 10.1167/iovs.64.10.16. Invest Ophthalmol Vis Sci. 2023. PMID: 37450309 Free PMC article.
-
Emerging Hypervirulent Marek's Disease Virus Variants Significantly Overcome Protection Conferred by Commercial Vaccines.Viruses. 2023 Jun 25;15(7):1434. doi: 10.3390/v15071434. Viruses. 2023. PMID: 37515122 Free PMC article.
-
High-Fidelity Long-Read Sequencing of an Avian Herpesvirus Reveals Extensive Intrapopulation Diversity in Tandem Repeat Regions.bioRxiv [Preprint]. 2025 Feb 11:2025.02.10.637388. doi: 10.1101/2025.02.10.637388. bioRxiv. 2025. PMID: 39990410 Free PMC article. Preprint.
References
-
- Read AF, Mackinnon MJ. Pathogen evolution in a vaccinated world In: Stearns SC, Koella JC, editors. Evolution in Health and Disease. 2nd ed Oxford, UK: Oxford University Press; 2007. p. 139–52.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

