Differential admixture in Latin American populations and its impact on the study of colorectal cancer
- PMID: 33306774
- PMCID: PMC7783724
- DOI: 10.1590/1678-4685-GMB-2020-0143
Differential admixture in Latin American populations and its impact on the study of colorectal cancer
Abstract
Genome-wide association studies focused on searching genes responsible for several diseases. Admixture mapping studies proposed a more efficient alternative capable of detecting polymorphisms contributing with a small effect on the disease risk. This method focuses on the higher values of linkage disequilibrium in admixed populations. To test this, we analyzed 10 genomic regions previously defined as related with colorectal cancer among nine populations and studied the variation pattern of haplotypic structures and heterozygosity values on seven categories of SNPs. Both analyses showed differences among chromosomal regions and studied populations. Admixed Latin-American samples generally show intermediate values. Heterozygosity of the SNPs grouped in categories varies more in each gene than in each population. African related populations have more blocks per chromosomal region, coherently with their antiquity. In sum, some similarities were found among Latin American populations, but each chromosomal region showed a particular behavior, despite the fact that the study refers to genes and regions related with one particular complex disease. This study strongly suggests the necessity of developing statistical methods to deal with di- or tri-hybrid populations, as well as to carefully analyze the different historic and demographic scenarios, and the different characteristics of particular chromosomal regions and evolutionary forces.
Figures


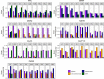
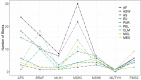
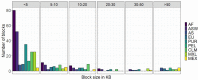
References
-
- Aaltonen LA, Johns L, Järvinen H, Mecklin JP, Houlston R. Explaining the familial colorectal cancer risk associated with mismatch repair (MMR)-deficient and MMR-stable tumors. Clin Cancer Res. 2007;13:356–361. - PubMed
-
- Botstein D, Risch N. Discovering genotypes underlying human phenotypes: Past successes for Mendelian disease, future approaches for complex disease. Nat Genet. 2003;33:228–237. - PubMed
LinkOut - more resources
Full Text Sources

