Prescribing Prevalence of Medications With Potential Genotype-Guided Dosing in Pediatric Patients
- PMID: 33315113
- PMCID: PMC7737091
- DOI: 10.1001/jamanetworkopen.2020.29411
Prescribing Prevalence of Medications With Potential Genotype-Guided Dosing in Pediatric Patients
Abstract
Importance: Genotype-guided prescribing in pediatrics could prevent adverse drug reactions and improve therapeutic response. Clinical pharmacogenetic implementation guidelines are available for many medications commonly prescribed to children. Frequencies of medication prescription and actionable genotypes (genotypes where a prescribing change may be indicated) inform the potential value of pharmacogenetic implementation.
Objective: To assess potential opportunities for genotype-guided prescribing in pediatric populations among multiple health systems by examining the prevalence of prescriptions for each drug with the highest level of evidence (Clinical Pharmacogenetics Implementation Consortium level A) and estimating the prevalence of potential actionable prescribing decisions.
Design, setting, and participants: This serial cross-sectional study of prescribing prevalences in 16 health systems included electronic health records data from pediatric inpatient and outpatient encounters from January 1, 2011, to December 31, 2017. The health systems included academic medical centers with free-standing children's hospitals and community hospitals that were part of an adult health care system. Participants included approximately 2.9 million patients younger than 21 years observed per year. Data were analyzed from June 5, 2018, to April 14, 2020.
Exposures: Prescription of 38 level A medications based on electronic health records.
Main outcomes and measures: Annual prevalence of level A medication prescribing and estimated actionable exposures, calculated by combining estimated site-year prevalences across sites with each site weighted equally.
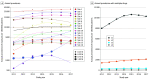
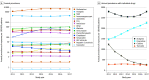
Results: Data from approximately 2.9 million pediatric patients (median age, 8 [interquartile range, 2-16] years; 50.7% female, 62.3% White) were analyzed for a typical calendar year. The annual prescribing prevalence of at least 1 level A drug ranged from 7987 to 10 629 per 100 000 patients with increasing trends from 2011 to 2014. The most prescribed level A drug was the antiemetic ondansetron (annual prevalence of exposure, 8107 [95% CI, 8077-8137] per 100 000 children). Among commonly prescribed opioids, annual prevalence per 100 000 patients was 295 (95% CI, 273-317) for tramadol, 571 (95% CI, 557-586) for codeine, and 2116 (95% CI, 2097-2135) for oxycodone. The antidepressants citalopram, escitalopram, and amitriptyline were also commonly prescribed (annual prevalence, approximately 250 per 100 000 patients for each). Estimated prevalences of actionable exposures were highest for oxycodone and ondansetron (>300 per 100 000 patients annually). CYP2D6 and CYP2C19 substrates were more frequently prescribed than medications influenced by other genes.
Conclusions and relevance: These findings suggest that opportunities for pharmacogenetic implementation among pediatric patients in the US are abundant. As expected, the greatest opportunity exists with implementing CYP2D6 and CYP2C19 pharmacogenetic guidance for commonly prescribed antiemetics, analgesics, and antidepressants.
Conflict of interest statement
Figures

References
-
- US Food and Drug Administration. Table of pharmacogenomic biomarkers in drug labeling. Updated August 18, 2020. Accessed March 16, 2020. https://www.fda.gov/drugs/science-and-research-drugs/table-pharmacogenom...
-
- US Food and Drug Administration Table of pharmacogenetic associations. Published February 25, 2020. Accessed March 31, 2020. https://www.fda.gov/medical-devices/precision-medicine/table-pharmacogen...
Publication types
MeSH terms
Substances
Grants and funding
- U01 HG007269/HG/NHGRI NIH HHS/United States
- U01 HG007253/HG/NHGRI NIH HHS/United States
- P30 CA076292/CA/NCI NIH HHS/United States
- K24 HL133373/HL/NHLBI NIH HHS/United States
- UL1 TR000165/TR/NCATS NIH HHS/United States
- UL1 TR002494/TR/NCATS NIH HHS/United States
- T32 HG008961/HG/NHGRI NIH HHS/United States
- U01 HG007775/HG/NHGRI NIH HHS/United States
- UL1 TR001427/TR/NCATS NIH HHS/United States
- K23 HD099331/HD/NICHD NIH HHS/United States
- R01 HL092173/HL/NHLBI NIH HHS/United States
- U01 HG010245/HG/NHGRI NIH HHS/United States
- U01 HG007762/HG/NHGRI NIH HHS/United States
- U01 HG010232/HG/NHGRI NIH HHS/United States
- T15 LM007450/LM/NLM NIH HHS/United States
- K23 HL143161/HL/NHLBI NIH HHS/United States
- U01 HG007278/HG/NHGRI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

