Advances in germline predisposition to acute leukaemias and myeloid neoplasms
- PMID: 33328584
- PMCID: PMC8404376
- DOI: 10.1038/s41568-020-00315-z
Advances in germline predisposition to acute leukaemias and myeloid neoplasms
Abstract
Although much work has focused on the elucidation of somatic alterations that drive the development of acute leukaemias and other haematopoietic diseases, it has become increasingly recognized that germline mutations are common in many of these neoplasms. In this Review, we highlight the different genetic pathways impacted by germline mutations that can ultimately lead to the development of familial and sporadic haematological malignancies, including acute lymphoblastic leukaemia, acute myeloid leukaemia (AML) and myelodysplastic syndrome (MDS). Many of the genes disrupted by somatic mutations in these diseases (for example, TP53, RUNX1, IKZF1 and ETV6) are the same as those that harbour germline mutations in children and adolescents who develop these malignancies. Moreover, the presumption that familial leukaemias only present in childhood is no longer true, in large part due to the numerous studies demonstrating germline DDX41 mutations in adults with MDS and AML. Lastly, we highlight how different cooperating events can influence the ultimate phenotype in these different familial leukaemia syndromes.
Figures


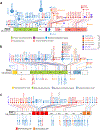

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

