Conserved Gsx2/Ind homeodomain monomer versus homodimer DNA binding defines regulatory outcomes in flies and mice
- PMID: 33334823
- PMCID: PMC7778271
- DOI: 10.1101/gad.343053.120
Conserved Gsx2/Ind homeodomain monomer versus homodimer DNA binding defines regulatory outcomes in flies and mice
Abstract
How homeodomain proteins gain sufficient specificity to control different cell fates has been a long-standing problem in developmental biology. The conserved Gsx homeodomain proteins regulate specific aspects of neural development in animals from flies to mammals, and yet they belong to a large transcription factor family that bind nearly identical DNA sequences in vitro. Here, we show that the mouse and fly Gsx factors unexpectedly gain DNA binding specificity by forming cooperative homodimers on precisely spaced and oriented DNA sites. High-resolution genomic binding assays revealed that Gsx2 binds both monomer and homodimer sites in the developing mouse ventral telencephalon. Importantly, reporter assays showed that Gsx2 mediates opposing outcomes in a DNA binding site-dependent manner: Monomer Gsx2 binding represses transcription, whereas homodimer binding stimulates gene expression. In Drosophila, the Gsx homolog, Ind, similarly represses or stimulates transcription in a site-dependent manner via an autoregulatory enhancer containing a combination of monomer and homodimer sites. Integrating these findings, we test a model showing how the homodimer to monomer site ratio and the Gsx protein levels defines gene up-regulation versus down-regulation. Altogether, these data serve as a new paradigm for how cooperative homeodomain transcription factor binding can increase target specificity and alter regulatory outcomes.
Keywords: CUT&RUN; Gsx2; Ind; lateral ganglionic eminence (LGE); transcriptional activation versus repression.
© 2021 Salomone et al.; Published by Cold Spring Harbor Laboratory Press.
Figures
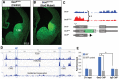
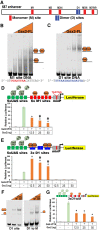



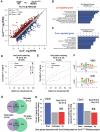


References
-
- Berger MF, Badis G, Gehrke AR, Talukder S, Philippakis AA, Peña-Castillo L, Alleyne TM, Mnaimneh S, Botvinnik OB, Chan ET, et al. 2008. Variation in homeodomain DNA binding revealed by high-resolution analysis of sequence preferences. Cell 133: 1266–1276. 10.1016/j.cell.2008.05.024 - DOI - PMC - PubMed
-
- Castro DS, Martynoga B, Parras C, Ramesh V, Pacary E, Johnston C, Drechsel D, Lebel-Potter M, Garcia LG, Hunt C, et al. 2011. A novel function of the proneural factor Ascl1 in progenitor proliferation identified by genome-wide characterization of its targets. Genes Dev 25: 930–945. 10.1101/gad.627811 - DOI - PMC - PubMed
-
- Corbin JG, Gaiano N, Machold RP, Langston A, Fishell G. 2000. The Gsh2 homeodomain gene controls multiple aspects of telencephalic development. Development 127: 5007–5020. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
