Identification of a six-gene metabolic signature predicting overall survival for patients with lung adenocarcinoma
- PMID: 33344071
- PMCID: PMC7718790
- DOI: 10.7717/peerj.10320
Identification of a six-gene metabolic signature predicting overall survival for patients with lung adenocarcinoma
Abstract
Background: Lung cancer is the leading cause of cancer-related deaths worldwide. Lung adenocarcinoma (LUAD) is one of the main subtypes of lung cancer. Hundreds of metabolic genes are altered consistently in LUAD; however, their prognostic role remains to be explored. This study aimed to establish a molecular signature that can predict the prognosis in patients with LUAD based on metabolic gene expression.
Methods: The transcriptome expression profiles and corresponding clinical information of LUAD were obtained from The Cancer Genome Atlas and Gene Expression Omnibus databases. The differentially expressed genes (DEGs) between LUAD and paired non-tumor samples were identified by the Wilcoxon rank sum test. Univariate Cox regression analysis and the lasso Cox regression model were used to construct the best-prognosis molecular signature. A nomogram was established comprising the prognostic model for predicting overall survival. To validate the prognostic ability of the molecular signature and the nomogram, the Kaplan-Meier survival analysis, Cox proportional hazards model, and receiver operating characteristic analysis were used.
Results: The six-gene molecular signature (PFKP, PKM, TPI1, LDHA, PTGES, and TYMS) from the DEGs was constructed to predict the prognosis. The molecular signature demonstrated a robust independent prognostic ability in the training and validation sets. The nomogram including the prognostic model had a greater predictive accuracy than previous systems. Furthermore, a gene set enrichment analysis revealed several significantly enriched metabolic pathways, which suggests a correlation of the molecular signature with metabolic systems and may help explain the underlying mechanisms.
Conclusions: Our study identified a novel six-gene metabolic signature for LUAD prognosis prediction. The molecular signature could reflect the dysregulated metabolic microenvironment, provide potential biomarkers for predicting prognosis, and indicate potential novel metabolic molecular-targeted therapies.
Keywords: GEO; Lung adenocarcinoma; Metabolic signature; Overall survival; Prognostic model; TCGA.
©2020 Cao et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures

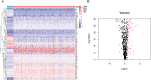








References
-
- Applebaum MA, Jha AR, Kao C, Hernandez KM, DeWane G, Salwen HR, Chlenski A, Dobratic M, Mariani CJ, Godley LA, Prabhakar N, White K, Stranger BE, Cohn SL. Integrative genomics reveals hypoxia inducible genes that are associated with a poor prognosis in neuroblastoma patients. Oncotarget. 2016;7(47):76816–76826. doi: 10.18632/oncotarget.12713. - DOI - PMC - PubMed
-
- Beer DG, Kardia SL, Huang CC, Giordano TJ, Levin AM, Misek DE, Lin L, Chen G, Gharib TG, Thomas DG, Lizyness ML, Kuick R, Hayasaka S, Taylor JM, Iannettoni MD, Orringer MB, Hanash S. Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nature Medicine. 2002;8(8):816–824. doi: 10.1038/nm733. - DOI - PubMed
-
- Bjerre MT, Strand SH, Nørgaard M, Kristensen H, Rasmussen AK, Mortensen MM, Fredsøe J, Mouritzen P, Ulhøi B, Ørntoft T, Borre M, Sørensen KD. Aberrant DOCK2, GRASP, HIF3A and PKFP hypermethylation has potential as a prognostic biomarker for prostate cancer. International Journal of Molecular Sciences. 2019;20(5):1173. doi: 10.3390/ijms20051173. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

