Assessment of transcriptional importance of cell line-specific features based on GTRD and FANTOM5 data
- PMID: 33347457
- PMCID: PMC7751965
- DOI: 10.1371/journal.pone.0243332
Assessment of transcriptional importance of cell line-specific features based on GTRD and FANTOM5 data
Abstract
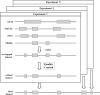
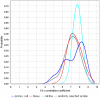
Creating a complete picture of the regulation of transcription seems to be an urgent task of modern biology. Regulation of transcription is a complex process carried out by transcription factors (TFs) and auxiliary proteins. Over the past decade, ChIP-Seq has become the most common experimental technology studying genome-wide interactions between TFs and DNA. We assessed the transcriptional significance of cell line-specific features using regression analysis of ChIP-Seq datasets from the GTRD database and transcriptional start site (TSS) activities from the FANTOM5 expression atlas. For this purpose, we initially generated a large number of features that were defined as the presence or absence of TFs in different promoter regions around TSSs. Using feature selection and regression analysis, we identified sets of the most important TFs that affect expression activity of TSSs in human cell lines such as HepG2, K562 and HEK293. We demonstrated that some TFs can be classified as repressors and activators depending on their location relative to TSS.
Conflict of interest statement
The authors have read the journal’s policy and have the following competing interests: RNS, YVK, ASR, ISY and FAK are employees of BIOSOFT.RU, Ltd. This does not alter our adherence to PLOS ONE policies on sharing data and materials. There are no patents, products in development or marketed products associated with this research to declare.
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous