Association of Vitamin D Metabolism Gene Polymorphisms with Autoimmunity: Evidence in Population Genetic Studies
- PMID: 33348854
- PMCID: PMC7766382
- DOI: 10.3390/ijms21249626
Association of Vitamin D Metabolism Gene Polymorphisms with Autoimmunity: Evidence in Population Genetic Studies
Abstract
A high prevalence of vitamin D (calcidiol) serum deficiency has been described in several autoimmune diseases, including multiple sclerosis (MS), rheumatoid arthritis (AR), and systemic lupus erythematosus (SLE). Vitamin D is a potent immunonutrient that through its main metabolite calcitriol, regulates the immunomodulation of macrophages, dendritic cells, T and B lymphocytes, which express the vitamin D receptor (VDR), and they produce and respond to calcitriol. Genetic association studies have shown that up to 65% of vitamin D serum variance may be explained due to genetic background. The 90% of genetic variability takes place in the form of single nucleotide polymorphisms (SNPs), and SNPs in genes related to vitamin D metabolism have been linked to influence the calcidiol serum levels, such as in the vitamin D binding protein (VDBP; rs2282679 GC), 25-hydroxylase (rs10751657 CYP2R1), 1α-hydroxylase (rs10877012, CYP27B1) and the vitamin D receptor (FokI (rs2228570), BsmI (rs1544410), ApaI (rs7975232), and TaqI (rs731236) VDR). Therefore, the aim of this comprehensive literature review was to discuss the current findings of functional SNPs in GC, CYP2R1, CYP27B1, and VDR associated to genetic risk, and the most common clinical features of MS, RA, and SLE.
Keywords: CYP27B1; CYP2R1; GC; VDR; autoimmune disease; vitamin D polymorphisms.
Conflict of interest statement
The authors declare no conflict of interest. Figures were created with BioRender software, ©biorender.com.
Figures
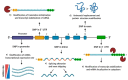

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

