Comparative genomics revealed adaptive admixture in Cryptosporidium hominis in Africa
- PMID: 33355530
- PMCID: PMC8115899
- DOI: 10.1099/mgen.0.000493
Comparative genomics revealed adaptive admixture in Cryptosporidium hominis in Africa
Abstract
Cryptosporidiosis is a major cause of diarrhoeal illness among African children, and is associated with childhood mortality, malnutrition, cognitive development and growth retardation. Cryptosporidium hominis is the dominant pathogen in Africa, and genotyping at the glycoprotein 60 (gp60) gene has revealed a complex distribution of different subtypes across this continent. However, a comprehensive exploration of the metapopulation structure and evolution based on whole-genome data has yet to be performed. Here, we sequenced and analysed the genomes of 26 C. hominis isolates, representing different gp60 subtypes, collected at rural sites in Gabon, Ghana, Madagascar and Tanzania. Phylogenetic and cluster analyses based on single-nucleotide polymorphisms showed that isolates predominantly clustered by their country of origin, irrespective of their gp60 subtype. We found a significant isolation-by-distance signature that shows the importance of local transmission, but we also detected evidence of hybridization between isolates of different geographical regions. We identified 37 outlier genes with exceptionally high nucleotide diversity, and this group is significantly enriched for genes encoding extracellular proteins and signal peptides. Furthermore, these genes are found more often than expected in recombinant regions, and they show a distinct signature of positive or balancing selection. We conclude that: (1) the metapopulation structure of C. hominis can only be accurately captured by whole-genome analyses; (2) local anthroponotic transmission underpins the spread of this pathogen in Africa; (3) hybridization occurs between distinct geographical lineages; and (4) genetic introgression provides novel substrate for positive or balancing selection in genes involved in host-parasite coevolution.
Keywords: Africa; Cryptosporidium hominis; genetic introgression; population structure; recombination; whole-genome sequencing.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures
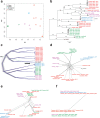


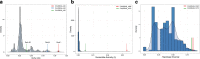
References
-
- Kotloff KL, Nataro JP, Blackwelder WC, Nasrin D, Farag TH, et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the global enteric multicenter study, GEMs): a prospective, case-control study. Lancet. 2013;382:209–222. - PubMed
-
- Molbak K, Andersen M, Aaby P, Hojlyng N, Jakobsen M, et al. Cryptosporidium infection in infancy as a cause of malnutrition: a community study from Guinea-Bissau, West Africa. Am J Clin Nutr. 1997;65:149–152. - PubMed
-
- Guerrant DI, Moore SR, Lima AA, Patrick PD, Schorling JB, et al. Association of early childhood diarrhea and cryptosporidiosis with impaired physical fitness and cognitive function four-seven years later in a poor urban community in northeast Brazil. Am J Trop Med Hyg. 1999;61:707–713. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

