Allosteric HIV Integrase Inhibitors Promote Formation of Inactive Branched Polymers via Homomeric Carboxy-Terminal Domain Interactions
- PMID: 33357410
- PMCID: PMC7935764
- DOI: 10.1016/j.str.2020.12.001
Allosteric HIV Integrase Inhibitors Promote Formation of Inactive Branched Polymers via Homomeric Carboxy-Terminal Domain Interactions
Abstract
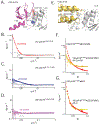
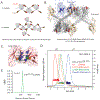
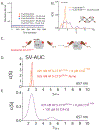
The major effect of allosteric HIV integrase (IN) inhibitors (ALLINIs) is observed during virion maturation, where ALLINI treatment interrupts IN-RNA interactions via drug-induced IN aggregation, leading to the formation of aberrant virions. To understand the structural changes that accompany drug-induced aggregation, we determined the soft matter properties of ALLINI-induced IN aggregates. Using small-angle neutron scattering, SEM, and rheology, we have discovered that the higher-order aggregates induced by ALLINIs have the characteristics of weak three-dimensional gels with a fractal-like character. Their formation is inhibited by the host factor LEDGF/p75, as well as ex vivo resistance substitutions. Mutagenesis and biophysical analyses reveal that homomeric carboxy-terminal domain interactions are required to achieve the branched-polymer nature of the ALLINI-induced aggregates. These studies provide key insight into the mechanisms of ALLINI action and resistance in the context of the crowded virion environment where ALLINIs exert their effect.
Keywords: ALLINI; HIV; analytical ultracentrifugation; host factors; oligomerization; polymer network; retroviral integration; rheology; small-angle X-ray scattering; small-angle neutron scattering.
Copyright © 2020 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare that they have no competing interests with the contents of this article.
Figures
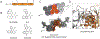



Similar articles
-
Structural Impact of Ex Vivo Resistance Mutations on HIV-1 Integrase Polymers Induced by Allosteric Inhibitors.J Mol Biol. 2025 Sep 1;437(17):169224. doi: 10.1016/j.jmb.2025.169224. Epub 2025 May 21. J Mol Biol. 2025. PMID: 40409709
-
The A128T resistance mutation reveals aberrant protein multimerization as the primary mechanism of action of allosteric HIV-1 integrase inhibitors.J Biol Chem. 2013 May 31;288(22):15813-20. doi: 10.1074/jbc.M112.443390. Epub 2013 Apr 24. J Biol Chem. 2013. PMID: 23615903 Free PMC article.
-
Allosteric integrase inhibitor potency is determined through the inhibition of HIV-1 particle maturation.Proc Natl Acad Sci U S A. 2013 May 21;110(21):8690-5. doi: 10.1073/pnas.1300703110. Epub 2013 Apr 22. Proc Natl Acad Sci U S A. 2013. PMID: 23610442 Free PMC article.
-
Multimodal mechanism of action of allosteric HIV-1 integrase inhibitors.Expert Rev Mol Med. 2013 Nov 26;15:e14. doi: 10.1017/erm.2013.15. Expert Rev Mol Med. 2013. PMID: 24274067 Free PMC article. Review.
-
Optimizing the Multimerization Properties of Quinoline-Based Allosteric HIV-1 Integrase Inhibitors.Viruses. 2024 Jan 28;16(2):200. doi: 10.3390/v16020200. Viruses. 2024. PMID: 38399977 Free PMC article. Review.
Cited by
-
Exploring Zinc C295 as a Dual HIV-1 Integrase Inhibitor: From Strand Transfer to 3'-Processing Suppression.Pharmaceuticals (Basel). 2024 Dec 29;18(1):30. doi: 10.3390/ph18010030. Pharmaceuticals (Basel). 2024. PMID: 39861093 Free PMC article.
-
Structure of a HIV-1 IN-Allosteric inhibitor complex at 2.93 Å resolution: Routes to inhibitor optimization.PLoS Pathog. 2023 Mar 3;19(3):e1011097. doi: 10.1371/journal.ppat.1011097. eCollection 2023 Mar. PLoS Pathog. 2023. PMID: 36867659 Free PMC article.
-
Brief Histories of Retroviral Integration Research and Associated International Conferences.Viruses. 2024 Apr 13;16(4):604. doi: 10.3390/v16040604. Viruses. 2024. PMID: 38675945 Free PMC article. Review.
-
The Drug-Induced Interface That Drives HIV-1 Integrase Hypermultimerization and Loss of Function.mBio. 2023 Feb 28;14(1):e0356022. doi: 10.1128/mbio.03560-22. Epub 2023 Feb 6. mBio. 2023. PMID: 36744954 Free PMC article.
-
Retrointegration2023-Papers from the 7th International Conference on Retroviral Integration.Viruses. 2025 Jun 23;17(7):879. doi: 10.3390/v17070879. Viruses. 2025. PMID: 40733497 Free PMC article.
References
-
- Allouch A, and Cereseto A (2011). Identification of cellular factors binding to acetylated HIV-1 integrase. Amino acids 41, 1137–1145. - PubMed
-
- Allouch A, Di Primio C, Alpi E, Lusic M, Arosio D, Giacca M, and Cereseto A (2011). The TRIM family protein KAP1 inhibits HIV-1 integration. Cell host & microbe 9, 484–495. - PubMed
-
- Bale HD, and Schmidt PW (1984). Small-Angle X-Ray-Scattering Investigation of Submicroscopic Porosity with Fractal Properties. Physical Review Letters 53, 596–599.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

