The Two-Component System CopRS Maintains Subfemtomolar Levels of Free Copper in the Periplasm of Pseudomonas aeruginosa Using a Phosphatase-Based Mechanism
- PMID: 33361129
- PMCID: PMC7763554
- DOI: 10.1128/mSphere.01193-20
The Two-Component System CopRS Maintains Subfemtomolar Levels of Free Copper in the Periplasm of Pseudomonas aeruginosa Using a Phosphatase-Based Mechanism
Abstract
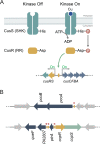
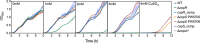
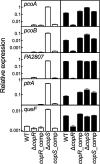
Two-component systems control periplasmic Cu+ homeostasis in Gram-negative bacteria. In characterized systems such as Escherichia coli CusRS, upon Cu+ binding to the periplasmic sensing region of CusS, a cytoplasmic phosphotransfer domain of the sensor phosphorylates the response regulator CusR. This drives the expression of efflux transporters, chaperones, and redox enzymes to ameliorate metal toxic effects. Here, we show that the Pseudomonas aeruginosa two-component sensor histidine kinase CopS exhibits a Cu-dependent phosphatase activity that maintains CopR in a nonphosphorylated state when the periplasmic Cu levels are below the activation threshold of CopS. Upon Cu+ binding to the sensor, the phosphatase activity is blocked and the phosphorylated CopR activates transcription of the CopRS regulon. Supporting the model, mutagenesis experiments revealed that the ΔcopS strain exhibits maximal expression of the CopRS regulon, lower intracellular Cu+ levels, and increased Cu tolerance compared to wild-type cells. The invariant phosphoacceptor residue His235 of CopS was not required for the phosphatase activity itself but was necessary for its Cu dependency. To sense the metal, the periplasmic domain of CopS binds two Cu+ ions at its dimeric interface. Homology modeling of CopS based on CusS structure (four Ag+ binding sites) clearly supports the different binding stoichiometries in the two systems. Interestingly, CopS binds Cu+/2+ with 3 × 10-14 M affinity, pointing to the absence of free (hydrated) Cu+/2+ in the periplasm.IMPORTANCE Copper is a micronutrient required as cofactor in redox enzymes. When free, copper is toxic, mismetallating proteins and generating damaging free radicals. Consequently, copper overload is a strategy that eukaryotic cells use to combat pathogens. Bacteria have developed copper-sensing transcription factors to control copper homeostasis. The cell envelope is the first compartment that has to cope with copper stress. Dedicated two-component systems control the periplasmic response to metal overload. This paper shows that the sensor kinase of the copper-sensing two-component system present in Pseudomonadales exhibits a signal-dependent phosphatase activity controlling the activation of its cognate response regulator, distinct from previously described periplasmic Cu sensors. Importantly, the data show that the system is activated by copper levels compatible with the absence of free copper in the cell periplasm. These observations emphasize the diversity of molecular mechanisms that have evolved in bacteria to manage the copper cellular distribution.
Keywords: Pseudomonas aeruginosa; copper; homeostasis; periplasm; two-component regulatory systems.
Copyright © 2020 Novoa-Aponte et al.
Figures
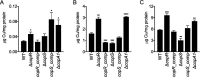



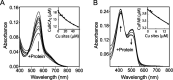

References
-
- Fraústo da Silva JJR, Williams RJP. 2001. The biological chemistry of the elements: the inorganic chemistry of life, 2nd ed Oxford University Press, Oxford, United Kingdom.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
