LSM12-EPAC1 defines a neuroprotective pathway that sustains the nucleocytoplasmic RAN gradient
- PMID: 33362237
- PMCID: PMC7757817
- DOI: 10.1371/journal.pbio.3001002
LSM12-EPAC1 defines a neuroprotective pathway that sustains the nucleocytoplasmic RAN gradient
Abstract
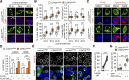
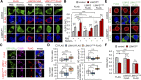
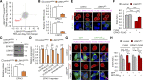
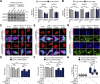
Nucleocytoplasmic transport (NCT) defects have been implicated in neurodegenerative diseases such as C9ORF72-associated amyotrophic lateral sclerosis and frontotemporal dementia (C9-ALS/FTD). Here, we identify a neuroprotective pathway of like-Sm protein 12 (LSM12) and exchange protein directly activated by cyclic AMP 1 (EPAC1) that sustains the nucleocytoplasmic RAN gradient and thereby suppresses NCT dysfunction by the C9ORF72-derived poly(glycine-arginine) protein. LSM12 depletion in human neuroblastoma cells aggravated poly(GR)-induced impairment of NCT and nuclear integrity while promoting the nuclear accumulation of poly(GR) granules. In fact, LSM12 posttranscriptionally up-regulated EPAC1 expression, whereas EPAC1 overexpression rescued the RAN gradient and NCT defects in LSM12-deleted cells. C9-ALS patient-derived neurons differentiated from induced pluripotent stem cells (C9-ALS iPSNs) displayed low expression of LSM12 and EPAC1. Lentiviral overexpression of LSM12 or EPAC1 indeed restored the RAN gradient, mitigated the pathogenic mislocalization of TDP-43, and suppressed caspase-3 activation for apoptosis in C9-ALS iPSNs. EPAC1 depletion biochemically dissociated RAN-importin β1 from the cytoplasmic nuclear pore complex, thereby dissipating the nucleocytoplasmic RAN gradient essential for NCT. These findings define the LSM12-EPAC1 pathway as an important suppressor of the NCT-related pathologies in C9-ALS/FTD.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Gijselinck I, Van Langenhove T, van der Zee J, Sleegers K, Philtjens S, Kleinberger G, et al. A C9orf72 promoter repeat expansion in a Flanders-Belgian cohort with disorders of the frontotemporal lobar degeneration-amyotrophic lateral sclerosis spectrum: a gene identification study. Lancet Neurol. 2012;11(1):54–65. 10.1016/S1474-4422(11)70261-7 - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

